Form 8-K Wave Life Sciences Ltd. For: May 13
Exhibit 99.1

Wave Life Sciences Reports First Quarter 2021 Financial Results and Provides Business Update
Clinical trials underway with next-generation candidates incorporating PN chemistry
Clinical data from PN chemistry programs expected in 2022
In vivo ADAR editing data for AATD program on track for 1H 2021
Wave to host investor conference call and webcast at 8:30 a.m. ET today
CAMBRIDGE, Mass., May 13, 2021 (GLOBE NEWSWIRE) — Wave Life Sciences Ltd. (Nasdaq: WVE), a clinical-stage genetic medicines company committed to delivering life-changing treatments for people battling devastating diseases, today announced financial results for the first quarter ended March 31, 2021 and provided a business update.
“Despite our PRECISION-HD results at the end of the first quarter, it has been a productive start of the year for Wave and our team remains focused on advancing our clinical trials for ALS/FTD, HD and DMD. These new trials mark the transition of our next-generation programs into the clinic. We expect clinical data that will provide insight into PN chemistry and enable decision making on next steps for these programs next year,” said Paul Bolno, MD, MBA, President and Chief Executive Officer of Wave Life Sciences. “We have a deep and diverse pipeline of RNA therapeutics, each designed with our PN chemistry, which has been shown to increase potency, exposure and durability compared to our first-generation compounds in preclinical studies. We continue to produce compelling in vivo data, and we are advancing multiple programs for CNS indications, including Alzheimer’s disease, Parkinson’s disease and others, in collaboration with our partner Takeda. Our ADAR editing capability demonstrates the diversity of our genetic medicines toolkit and we are well-positioned to be leaders in the RNA editing field. We look forward to providing more updates on ADAR editing, including the first in vivo data from our AATD program, in the first half of this year.”
Recent Business Highlights and Upcoming Milestones
WVE-004 (C9orf72) for amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD):
| • | WVE-004 is an investigational antisense oligonucleotide designed to selectively target transcript variants containing a hexanucleotide repeat expansion (G4C2) in the C9orf72 gene, which is one of the most common genetic causes of the sporadic and inherited forms of ALS and FTD. WVE-004 uses novel PN backbone chemistry modifications. |
| • | In February 2021, Wave published in Nature Communications the results of initial work to identify and validate its targeting strategy to achieve variant-selective knockdown of expansion-containing C9orf72 transcripts. |
| • | In April 2021, during a platform presentation at the American Academy of Neurology (AAN) 2021 Virtual Annual Meeting, Wave highlighted preclinical in vivo data for WVE-004, which demonstrated potent and durable knockdown of more than 90% of polyGP dipeptide repeat (DPR) proteins in the spinal cord and at least 80% in the cortex, an effect that persisted for at least six months. C9orf72 protein was relatively unchanged over the same time period. |
| • | This week, at the European Network to Cure ALS (ENCALS) meeting being held May 12 – May 14, Wave is presenting a poster introducing its FOCUS-C9 Phase 1b/2a trial design for WVE-004. The FOCUS-C9 trial is a global, multicenter, randomized, double-blind, placebo-controlled Phase 1b/2a clinical trial to assess the safety and tolerability of intrathecal doses of WVE-004 for patients with C9-ALS and/or C9-FTD. Additional objectives include measurement of polyGP proteins in the cerebrospinal fluid (CSF), plasma and CSF pharmacokinetics, and exploratory biomarker and clinical endpoints. The FOCUS-C9 trial is designed to be adaptive and includes single- and multiple-ascending dose portions, with dose escalation and dosing frequency being guided by an independent safety committee. |
| • | Wave has received regulatory and ethics approvals and site activation is underway for the FOCUS-C9 clinical trial, and Wave expects to initiate dosing in 2021. |
WVE-003 (SNP3) for Huntington’s disease (HD):
| • | WVE-003 is Wave’s next-generation HD candidate and Wave’s first HD candidate that uses PN chemistry. WVE-003 is designed to selectively target the mutant allele of the huntingtin (mHTT) gene, while leaving the wild-type (wtHTT) protein relatively intact. Wave’s approach to HD is guided by the recognition that, in addition to a gain of function of the mHTT protein, people with this disease have less wtHTT protein, leaving them with a smaller protective reservoir of healthy protein than unaffected individuals. A growing body of scientific evidence suggests that preserving as much of this essential protein as possible, when in the setting of stress from toxic mHTT protein, may be important for favorable clinical outcomes. |
| • | In April 2021, at the 16th Annual CHDI Foundation Huntington’s Disease Therapeutic Conference, Wave highlighted preclinical data for WVE-003, which showed selective reduction of mHTT mRNA in vitro and potent and durable knockdown of mHTT mRNA in vivo. Wave also introduced the design for the Phase 1b/2a clinical trial of WVE-003, called SELECT-HD. The multicenter, randomized, double-blind, placebo-controlled trial will assess the safety and tolerability of intrathecally administered WVE-003 for patients with early manifest Huntington’s disease. Additional objectives include measurement of mHTT and wtHTT protein and exploratory pharmacokinetic, pharmacodynamic, clinical and MRI endpoints. The trial is designed to be adaptive, with dose escalation and dosing frequency being guided by an independent safety committee. |
| • | Wave has received regulatory and ethics approvals and site activation is underway for the SELECT-HD clinical trial, and Wave expects to initiate dosing in 2021. |
WVE-N531 for Duchenne muscular dystrophy (DMD) amenable to exon 53 skipping:
| • | WVE-N531 is Wave’s first splicing candidate to incorporate PN chemistry, which Wave advanced following results of an in vivo study in double knock-out mice (dKO) that showed that an oligonucleotide designed with PN chemistry appeared to significantly increase dystrophin production and substantially improve survival, compared to oligonucleotides designed with Wave’s first-generation chemistry. |
| • | In March 2021, Wave initiated clinical development of WVE-N531 with the submission of a clinical trial application. |
| • | Wave has received regulatory approval for a clinical trial of WVE-N531 to assess initial safety and dystrophin production in patients with DMD amenable to exon 53 skipping. Wave expects to initiate dosing in this trial in 2021. |
ADAR editing:
| • | In March 2021, Wave presented a poster at the 2021 Keystone eSymposia on Precision Engineering of the Genome, Epigenome and Transcriptome highlighting the breadth of RNA editing data generated using its ADAR editing capability to date. This presentation illustrated editing activity across in vivo and in vitro systems, including in vivo editing in the CNS, using conjugated and non-conjugated oligonucleotides. Wave will also present these data in an oral presentation at the 24th American Society of Gene and Cell Therapy (ASGCT) Annual Meeting being held this week, May 11 – 14, 2021. |
| • | Wave expects to present additional ADAR editing data at scientific congresses in 2021. |
Alpha-1 antitrypsin deficiency (AATD) program with ADAR editing:
| • | Wave’s AATD program, its first ADAR editing program, uses an oligonucleotide to correct the single RNA base mutation in mRNA coded by the SERPINA1 Z allele. ADAR editing may provide an ideal approach to treating AATD by increasing circulating levels of healthy alpha-1 antitrypsin (AAT) protein and reducing aggregation in the liver, thus simultaneously addressing both the lung and liver manifestations of the disease. |
| • | To support the continued development of its AATD program, Wave has developed a proprietary humanized SERPINA1/ADAR model. Wave expects to share in vivo data from this model in the first half of 2021 and plans to submit these data for presentation at a scientific congress in 2021. |
First Quarter 2021 Financial Results and Financial Guidance
Wave reported a net loss of $42.5 million in the first quarter of 2021 as compared to $47.5 million in the same period in 2020.
Research and development expenses were $33.4 million in the first quarter of 2021 as compared to $41.2 million in the same period in 2020. The year-over-year decrease was primarily due to the decrease in external expenses related to Wave’s suvodirsen program, which was discontinued in December 2019, but had wind-down costs throughout 2020, as well as decreases in compensation-related expenses and other external expenses, partially offset by the increases in external expenses related to Wave’s clinical and preclinical activities related to its HD programs and its C9orf72 program for ALS and FTD.
General and administrative expenses were $10.1 million in the first quarter of 2021, as compared to $13.0 million in the same period in 2020. The year-over-year decrease was driven by decreases in compensation-related expenses and other external expenses.
Wave ended the first quarter of 2021 with $148.5 million in cash and cash equivalents, as compared to $184.5 million as of December 31, 2020. The decrease in cash and cash equivalents was mainly due to Wave’s year-to-date net loss, partially offset by the receipt of $8.0 million in net proceeds under Wave’s at-the-market equity program. In April 2021, Wave received an additional $30.0 million in committed research support under its collaboration with Takeda.
Wave expects that its existing cash and cash equivalents, together with expected and committed cash from its existing collaboration, will enable the company to fund its operating and capital expenditure requirements into the second quarter of 2023.
Investor Conference Call and Webcast
Wave management will host an investor conference call today at 8:30 a.m. ET to discuss the company’s first quarter and 2021 financial results and provide a business update. The conference call may be accessed by dialing (866) 220-8068 (domestic) or (470) 495-9153 (international) and entering conference ID: 7430859. The live webcast may be accessed from the investor relations section of the Wave Life Sciences corporate website at ir.wavelifesciences.com. Following the webcast, a replay will be available on the website.
About PRISM™
PRISM is Wave Life Sciences’ proprietary discovery and drug development platform that enables genetically defined diseases to be targeted with stereopure oligonucleotides across multiple therapeutic modalities, including silencing, splicing and editing. PRISM combines the company’s unique ability to construct stereopure oligonucleotides with a deep understanding of how the interplay among oligonucleotide sequence, chemistry and backbone stereochemistry impacts key pharmacological properties. By exploring these interactions through iterative analysis of in vitro and in vivo outcomes and machine learning-driven predictive modeling, the company continues to define design principles that are deployed across programs to rapidly develop and manufacture clinical candidates that meet pre-defined product profiles.
About Wave Life Sciences
Wave Life Sciences (Nasdaq: WVE) is a clinical-stage genetic medicines company committed to delivering life-changing treatments for people battling devastating diseases. Wave aspires to develop best-in-class medicines across multiple therapeutic modalities using PRISM, the company’s proprietary discovery and drug development platform that enables the precise design, optimization and production of stereopure oligonucleotides. Driven by a resolute sense of urgency, the Wave team is targeting a broad range of genetically defined diseases so that patients and families may realize a brighter future. To find out more, please visit www.wavelifesciences.com and follow Wave on Twitter @WaveLifeSci.
Forward-Looking Statements
This press release contains forward-looking statements concerning our goals, beliefs, expectations, strategies, objectives and plans, and other statements that are not necessarily based on historical facts, including statements regarding the following, among others: the anticipated commencement, patient enrollment, data readouts and completion of our adaptive clinical trials, and the announcement of such events; the protocol, design and endpoints of our ongoing and planned clinical trials; the future performance and results of our programs in clinical trials; future preclinical activities and programs; regulatory submissions; the progress and potential benefits of our collaborations with partners; the potential of our in vitro and in vivo preclinical data to predict the behavior of our compounds in humans; our identification of future product candidates and their therapeutic potential; the anticipated therapeutic benefits of our potential therapies compared to others; our ability to design compounds using multiple modalities and the anticipated benefits of that model; the anticipated benefits of our proprietary manufacturing processes and our internal manufacturing capabilities; the potential benefits of PRISM, including our novel PN backbone chemistry modifications, and our stereopure oligonucleotides compared with stereorandom oligonucleotides; the potential benefits of our novel ADAR-mediated RNA editing platform capabilities compared to others; the benefit of nucleic acid therapeutics generally; the strength of our intellectual property; the anticipated duration of our cash runway; and our expectations regarding the impact of the COVID-19 pandemic on our business. Actual results may differ materially from those indicated by these forward-looking statements as a result of various important factors, including the following: our ability to finance our drug discovery and development efforts and to raise additional capital when needed; the ability of our preclinical programs to produce data sufficient to support our clinical trial applications and the timing thereof; our ability to maintain the company infrastructure and personnel needed to achieve our goals; the clinical results of our programs, which may not support further development of product candidates; actions of regulatory agencies, which may affect the initiation, timing and progress of clinical trials, including their receptiveness to our adaptive trial designs; our effectiveness in managing future clinical trials and regulatory interactions; the effectiveness of PRISM, including our novel PN backbone chemistry modifications ; the effectiveness of our novel ADAR-mediated RNA editing platform capability; the continued development and acceptance of oligonucleotides as a class of medicines; our ability to demonstrate the therapeutic benefits of our candidates in clinical trials, including our ability to develop candidates across multiple therapeutic modalities; our dependence on third parties, including contract research organizations, contract manufacturing organizations, collaborators and partners; our ability to manufacture or contract with third parties to manufacture drug material to support our programs and growth; our ability to obtain, maintain and protect our intellectual property; our ability to enforce our patents against infringers and defend our patent portfolio against challenges from third parties; competition from others developing therapies for similar indications; the severity and duration of the COVID-19 pandemic and its negative impact on the conduct of, and the timing of enrollment, completion and reporting with respect to, our clinical trials; and any other impacts on our business as a result of or related to the COVID-19 pandemic, as well as the information under the caption “Risk Factors” contained in our most recent Annual Report on Form 10-K filed with the Securities and Exchange Commission (SEC) and in other filings we make with the SEC from time to time. We undertake no obligation to update the information contained in this press release to reflect subsequently occurring events or circumstances.
WAVE LIFE SCIENCES LTD.
UNAUDITED CONSOLIDATED BALANCE SHEETS
(In thousands, except share amounts)
| March 31, 2021 | December 31, 2020 | |||||||
| Assets |
||||||||
| Current assets: |
||||||||
| Cash and cash equivalents |
$ | 148,535 | $ | 184,497 | ||||
| Current portion of accounts receivable |
30,000 | 30,000 | ||||||
| Prepaid expenses |
10,430 | 10,434 | ||||||
| Other current assets |
5,580 | 5,111 | ||||||
|
|
|
|
|
|||||
| Total current assets |
194,545 | 230,042 | ||||||
|
|
|
|
|
|||||
| Long-term assets: |
||||||||
| Property and equipment, net |
27,370 | 29,198 | ||||||
| Operating lease right-of-use assets |
15,720 | 16,232 | ||||||
| Restricted cash |
3,651 | 3,651 | ||||||
| Other assets |
1,361 | 115 | ||||||
|
|
|
|
|
|||||
| Total long-term assets |
48,102 | 49,196 | ||||||
|
|
|
|
|
|||||
| Total assets |
$ | 242,647 | $ | 279,238 | ||||
|
|
|
|
|
|||||
| Liabilities, Series A preferred shares and shareholders’ equity |
||||||||
| Current liabilities: |
||||||||
| Accounts payable |
$ | 13,418 | $ | 13,795 | ||||
| Accrued expenses and other current liabilities |
6,661 | 11,971 | ||||||
| Current portion of deferred revenue |
24,763 | 91,560 | ||||||
| Current portion of operating lease liability |
3,838 | 3,714 | ||||||
|
|
|
|
|
|||||
| Total current liabilities |
48,680 | 121,040 | ||||||
|
|
|
|
|
|||||
| Long-term liabilities: |
||||||||
| Deferred revenue, net of current portion |
108,278 | 41,481 | ||||||
| Operating lease liability, net of current portion |
24,587 | 25,591 | ||||||
| Other liabilities |
407 | 474 | ||||||
|
|
|
|
|
|||||
| Total long-term liabilities |
133,272 | 67,546 | ||||||
|
|
|
|
|
|||||
| Total liabilities |
$ | 181,952 | $ | 188,586 | ||||
|
|
|
|
|
|||||
| Series A preferred shares, no par value; 3,901,348 shares issued and outstanding at March 31, 2021 and December 31, 2020 |
$ | 7,874 | $ | 7,874 | ||||
|
|
|
|
|
|||||
| Shareholders’ equity: |
||||||||
| Ordinary shares, no par value; 49,854,651 and 48,778,678 shares issued and outstanding at March 31, 2021 and December 31, 2020, respectively |
$ | 702,649 | $ | 694,085 | ||||
| Additional paid-in capital |
75,636 | 71,573 | ||||||
| Accumulated other comprehensive income |
269 | 389 | ||||||
| Accumulated deficit |
(725,733 | ) | (683,269 | ) | ||||
|
|
|
|
|
|||||
| Total shareholders’ equity |
$ | 52,821 | $ | 82,778 | ||||
|
|
|
|
|
|||||
| Total liabilities, Series A preferred shares and shareholders’ equity |
$ | 242,647 | $ | 279,238 | ||||
|
|
|
|
|
|||||
WAVE LIFE SCIENCES LTD.
UNAUDITED CONSOLIDATED STATEMENTS OF OPERATIONS AND COMPREHENSIVE LOSS
(In thousands, except share and per share amounts)
| Three Months Ended March 31, | ||||||||
| 2021 | 2020 | |||||||
| Revenue |
$ | — | $ | 4,161 | ||||
|
|
|
|
|
|||||
| Operating expenses: |
||||||||
| Research and development |
33,393 | 41,158 | ||||||
| General and administrative |
10,078 | 12,996 | ||||||
|
|
|
|
|
|||||
| Total operating expenses |
43,471 | 54,154 | ||||||
|
|
|
|
|
|||||
| Loss from operations |
(43,471 | ) | (49,993 | ) | ||||
| Other income, net: |
||||||||
| Dividend income and interest income, net |
11 | 388 | ||||||
| Other income, net |
996 | 2,112 | ||||||
|
|
|
|
|
|||||
| Total other income, net |
1,007 | 2,500 | ||||||
|
|
|
|
|
|||||
| Loss before income taxes |
(42,464 | ) | (47,493 | ) | ||||
| Income tax provision |
— | — | ||||||
|
|
|
|
|
|||||
| Net loss |
$ | (42,464 | ) | $ | (47,493 | ) | ||
|
|
|
|
|
|||||
| Net loss per share attributable to ordinary shareholders—basic and diluted |
$ | (0.86 | ) | $ | (1.38 | ) | ||
|
|
|
|
|
|||||
| Weighted-average ordinary shares used in computing net loss per share attributable to ordinary shareholders—basic and diluted |
49,101,606 | 34,461,505 | ||||||
|
|
|
|
|
|||||
| Other comprehensive income (loss): |
||||||||
| Net loss |
$ | (42,464 | ) | $ | (47,493 | ) | ||
| Foreign currency translation |
(120 | ) | 6 | |||||
|
|
|
|
|
|||||
| Comprehensive loss |
$ | (42,584 | ) | $ | (47,487 | ) | ||
|
|
|
|
|
|||||
Investor Contact:
Kate Rausch
617-949-4827
Media Contact:
Alicia Suter
617-949-4817

Wave Life Sciences Corporate Presentation May 13, 2021 Exhibit 99.2

Forward-looking statements This document contains forward-looking statements. All statements other than statements of historical facts contained in this document, including statements regarding possible or assumed future results of operations, preclinical and clinical studies, business strategies, research and development plans, collaborations and partnerships, regulatory activities and timing thereof, competitive position, potential growth opportunities, use of proceeds and the effects of competition are forward-looking statements. These statements involve known and unknown risks, uncertainties and other important factors that may cause the actual results, performance or achievements of Wave Life Sciences Ltd. (the “Company”) to be materially different from any future results, performance or achievements expressed or implied by the forward-looking statements. In some cases, you can identify forward-looking statements by terms such as “may,” “will,” “should,” “expect,” “plan,” “aim,” “anticipate,” “could,” “intend,” “target,” “project,” “contemplate,” “believe,” “estimate,” “predict,” “potential” or “continue” or the negative of these terms or other similar expressions. The forward-looking statements in this presentation are only predictions. The Company has based these forward-looking statements largely on its current expectations and projections about future events and financial trends that it believes may affect the Company’s business, financial condition and results of operations. These forward-looking statements speak only as of the date of this presentation and are subject to a number of risks, uncertainties and assumptions, including those listed under Risk Factors in the Company’s Form 10-K and other filings with the SEC, some of which cannot be predicted or quantified and some of which are beyond the Company’s control. The events and circumstances reflected in the Company’s forward-looking statements may not be achieved or occur, and actual results could differ materially from those projected in the forward-looking statements. Moreover, the Company operates in a dynamic industry and economy. New risk factors and uncertainties may emerge from time to time, and it is not possible for management to predict all risk factors and uncertainties that the Company may face. Except as required by applicable law, the Company does not plan to publicly update or revise any forward-looking statements contained herein, whether as a result of any new information, future events, changed circumstances or otherwise.
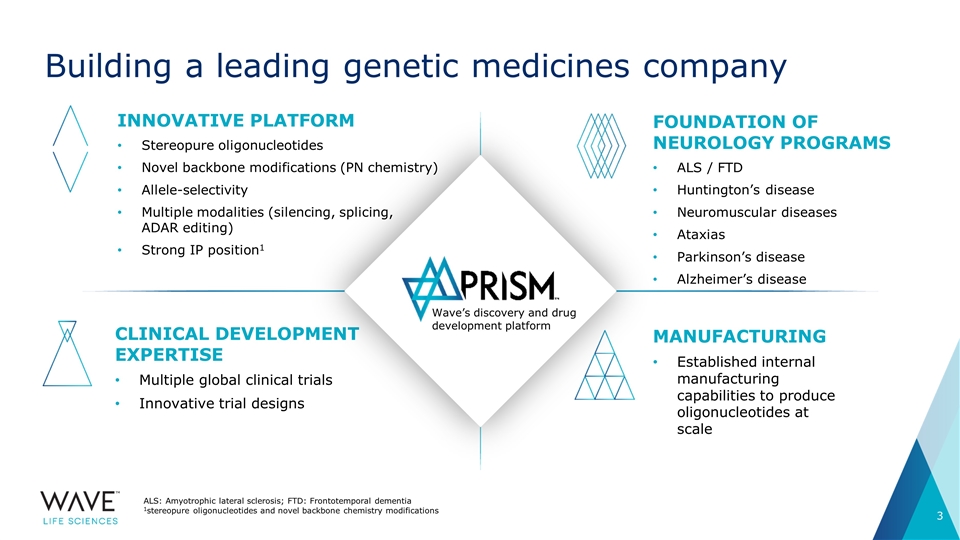
Building a leading genetic medicines company ALS: Amyotrophic lateral sclerosis; FTD: Frontotemporal dementia 1stereopure oligonucleotides and novel backbone chemistry modifications Innovative platform Stereopure oligonucleotides Novel backbone modifications (PN chemistry) Allele-selectivity Multiple modalities (silencing, splicing, ADAR editing) Strong IP position1 Foundation of NEUROLOGY programs ALS / FTD Huntington’s disease Neuromuscular diseases Ataxias Parkinson’s disease Alzheimer’s disease Clinical development expertise Multiple global clinical trials Innovative trial designs Manufacturing Established internal manufacturing capabilities to produce oligonucleotides at scale Wave’s discovery and drug development platform
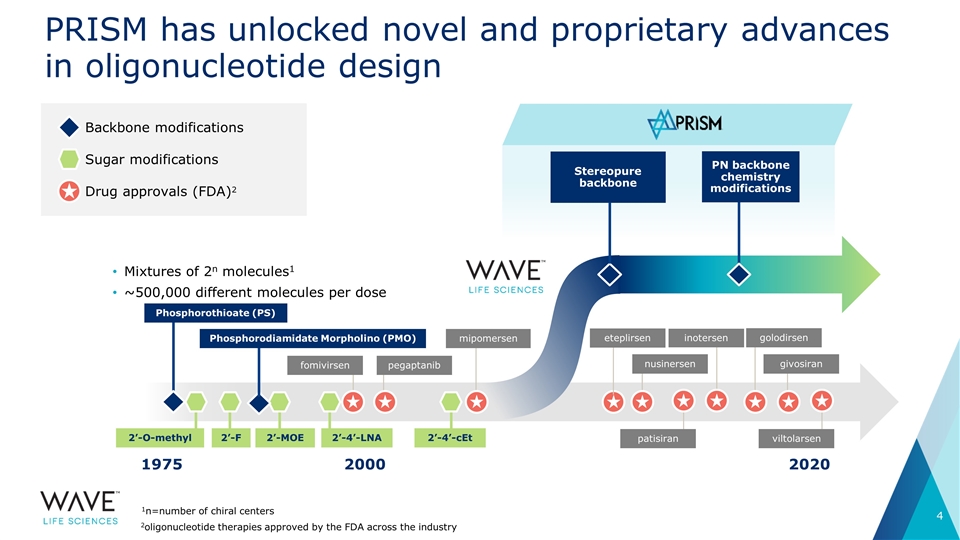
PRISM has unlocked novel and proprietary advances in oligonucleotide design Backbone modifications Sugar modifications Drug approvals (FDA)2 1975 2020 2000 Mixtures of 2n molecules1 ~500,000 different molecules per dose fomivirsen pegaptanib Phosphorothioate (PS) mipomersen nusinersen PN backbone chemistry modifications Stereopure backbone 2’-4’-cEt 2’-O-methyl 2’-F 2’-4’-LNA 1n=number of chiral centers 2’-MOE Phosphorodiamidate Morpholino (PMO) eteplirsen golodirsen givosiran patisiran inotersen viltolarsen 2oligonucleotide therapies approved by the FDA across the industry
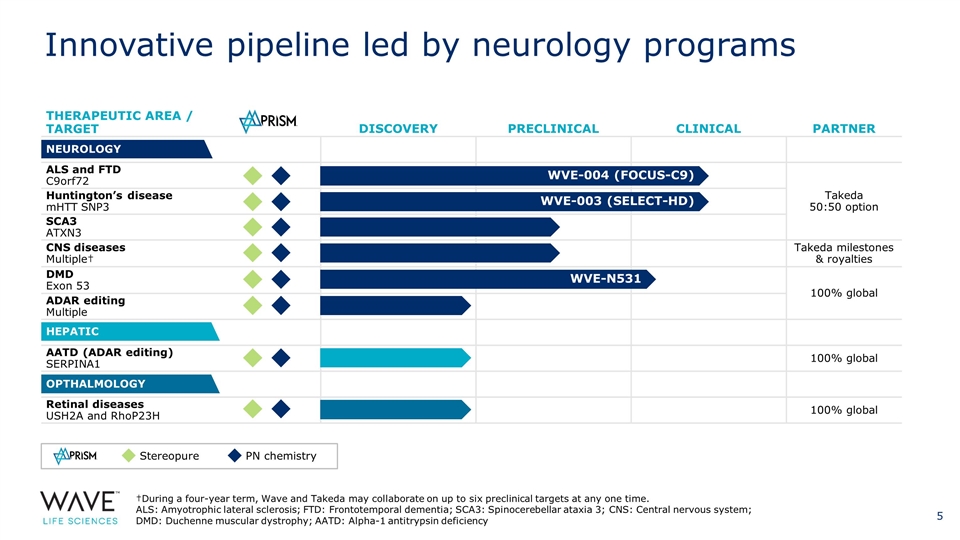
THERAPEUTIC AREA / TARGET DISCOVERY PRECLINICAL CLINICAL PARTNER ALS and FTD C9orf72 Takeda 50:50 option Huntington’s disease mHTT SNP3 SCA3 ATXN3 CNS diseases Multiple† Takeda milestones & royalties DMD Exon 53 100% global ADAR editing Multiple AATD (ADAR editing) SERPINA1 100% global Retinal diseases USH2A and RhoP23H 100% global NEUROLOGY HEPATIC OPTHALMOLOGY WVE-004 (FOCUS-C9) WVE-003 (SELECT-HD) †During a four-year term, Wave and Takeda may collaborate on up to six preclinical targets at any one time. ALS: Amyotrophic lateral sclerosis; FTD: Frontotemporal dementia; SCA3: Spinocerebellar ataxia 3; CNS: Central nervous system; DMD: Duchenne muscular dystrophy; AATD: Alpha-1 antitrypsin deficiency Stereopure PN chemistry WVE-N531 Innovative pipeline led by neurology programs
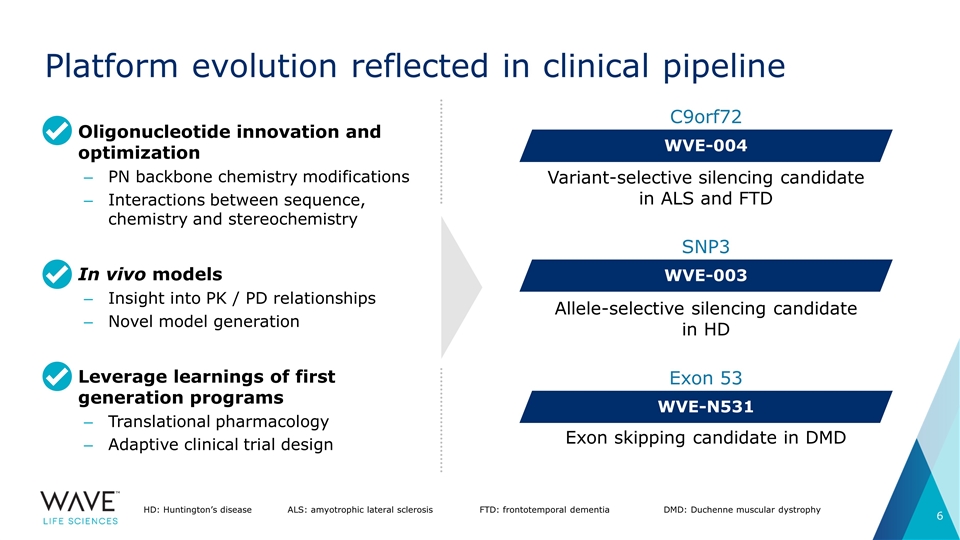
Platform evolution reflected in clinical pipeline Variant-selective silencing candidate in ALS and FTD WVE-004 C9orf72 WVE-003 SNP3 Allele-selective silencing candidate in HD WVE-N531 Exon 53 Exon skipping candidate in DMD HD: Huntington’s diseaseALS: amyotrophic lateral sclerosisFTD: frontotemporal dementia DMD: Duchenne muscular dystrophy Oligonucleotide innovation and optimization PN backbone chemistry modifications Interactions between sequence, chemistry and stereochemistry In vivo models Insight into PK / PD relationships Novel model generation Leverage learnings of first generation programs Translational pharmacology Adaptive clinical trial design

WVE-004 Amyotrophic Lateral Sclerosis (ALS) Frontotemporal Dementia (FTD)
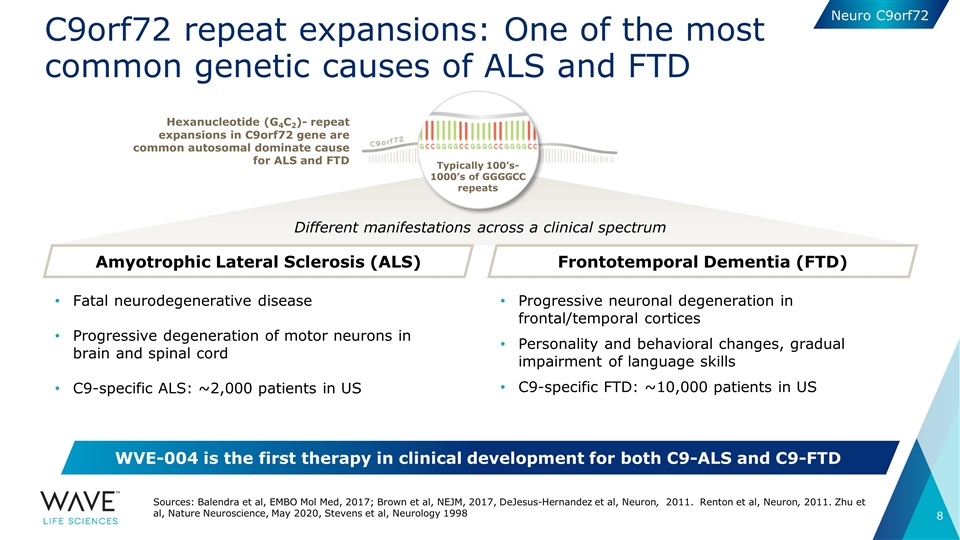
C9orf72 repeat expansions: One of the most common genetic causes of ALS and FTD Typically 100’s-1000’s of GGGGCC repeats Amyotrophic Lateral Sclerosis (ALS) Frontotemporal Dementia (FTD) Hexanucleotide (G4C2)- repeat expansions in C9orf72 gene are common autosomal dominate cause for ALS and FTD Different manifestations across a clinical spectrum Fatal neurodegenerative disease Progressive degeneration of motor neurons in brain and spinal cord C9-specific ALS: ~2,000 patients in US Progressive neuronal degeneration in frontal/temporal cortices Personality and behavioral changes, gradual impairment of language skills C9-specific FTD: ~10,000 patients in US WVE-004 is the first therapy in clinical development for both C9-ALS and C9-FTD Sources: Balendra et al, EMBO Mol Med, 2017; Brown et al, NEJM, 2017, DeJesus-Hernandez et al, Neuron, 2011. Renton et al, Neuron, 2011. Zhu et al, Nature Neuroscience, May 2020, Stevens et al, Neurology 1998 Neuro C9orf72
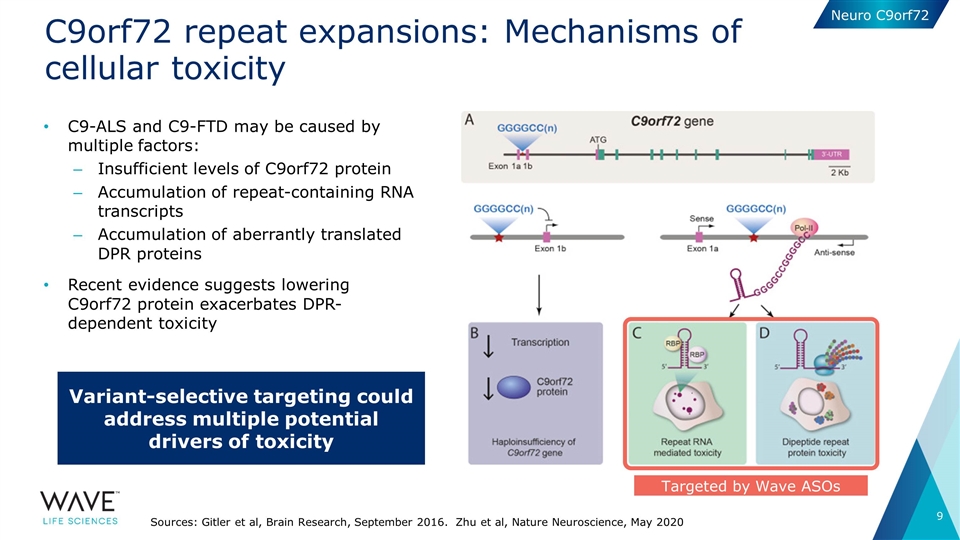
C9orf72 repeat expansions: Mechanisms of cellular toxicity C9-ALS and C9-FTD may be caused by multiple factors: Insufficient levels of C9orf72 protein Accumulation of repeat-containing RNA transcripts Accumulation of aberrantly translated DPR proteins Recent evidence suggests lowering C9orf72 protein exacerbates DPR-dependent toxicity Sources: Gitler et al, Brain Research, September 2016. Zhu et al, Nature Neuroscience, May 2020 Targeted by Wave ASOs Variant-selective targeting could address multiple potential drivers of toxicity Neuro C9orf72
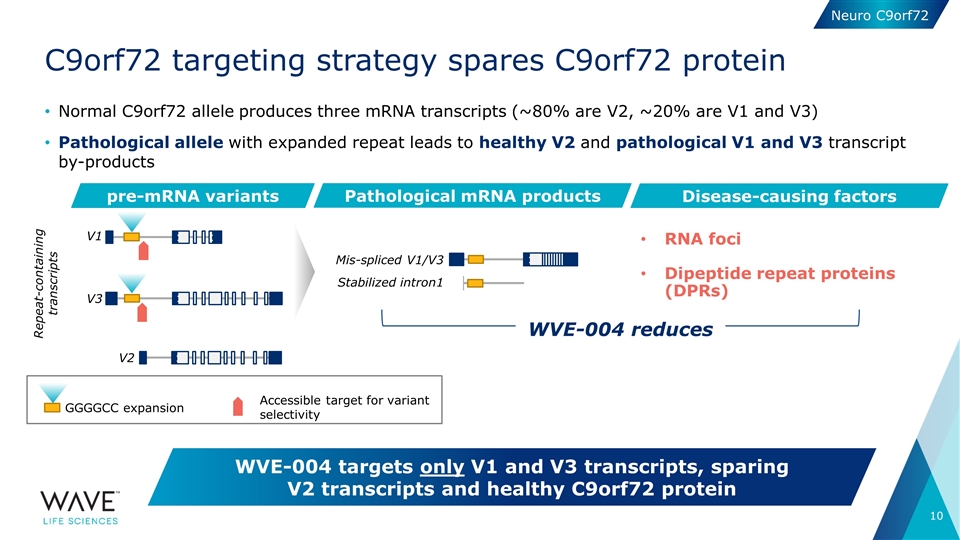
Normal C9orf72 allele produces three mRNA transcripts (~80% are V2, ~20% are V1 and V3) Pathological allele with expanded repeat leads to healthy V2 and pathological V1 and V3 transcript by-products C9orf72 targeting strategy spares C9orf72 protein WVE-004 targets only V1 and V3 transcripts, sparing V2 transcripts and healthy C9orf72 protein pre-mRNA variants Pathological mRNA products V1 V2 Mis-spliced V1/V3 Stabilized intron1 V3 Disease-causing factors RNA foci Dipeptide repeat proteins (DPRs) GGGGCC expansion Accessible target for variant selectivity WVE-004 reduces Repeat-containing transcripts Neuro C9orf72
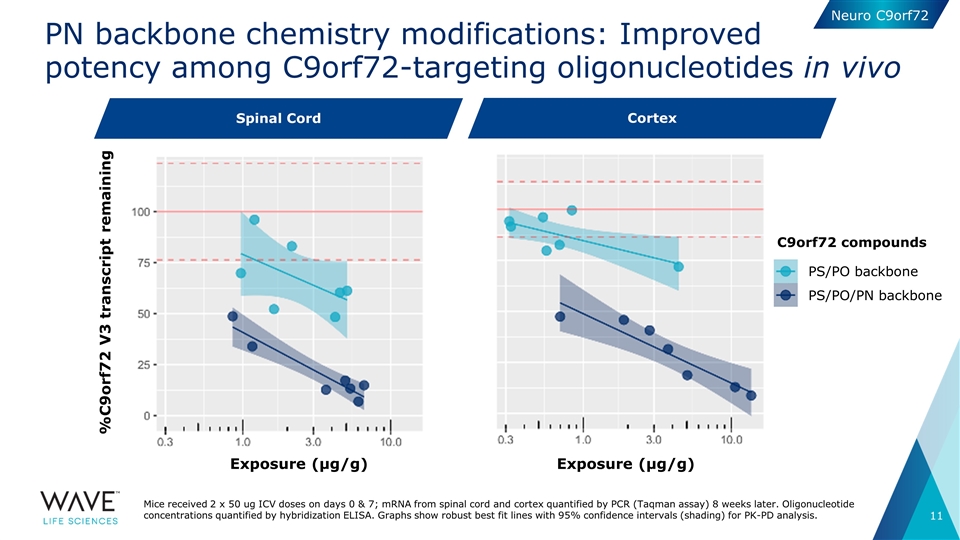
PN backbone chemistry modifications: Improved potency among C9orf72-targeting oligonucleotides in vivo Exposure (µg/g) Exposure (µg/g) C9orf72 compounds Spinal cord Cortex PS/PO backbone PS/PO/PN backbone %C9orf72 V3 transcript remaining Mice received 2 x 50 ug ICV doses on days 0 & 7; mRNA from spinal cord and cortex quantified by PCR (Taqman assay) 8 weeks later. Oligonucleotide concentrations quantified by hybridization ELISA. Graphs show robust best fit lines with 95% confidence intervals (shading) for PK-PD analysis. Spinal Cord Neuro C9orf72
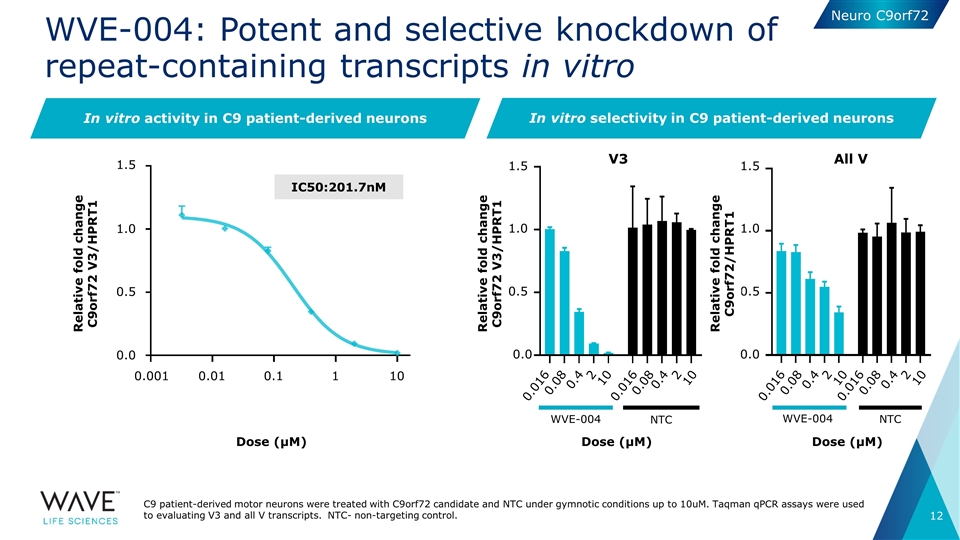
WVE-004: Potent and selective knockdown of repeat-containing transcripts in vitro V3 Dose (μM) All V WVE-004 NTC Dose (μM) In vitro activity in C9 patient-derived neurons WVE-004 NTC Dose (μM) IC50:201.7nM In vitro selectivity in C9 patient-derived neurons C9 patient-derived motor neurons were treated with C9orf72 candidate and NTC under gymnotic conditions up to 10uM. Taqman qPCR assays were used to evaluating V3 and all V transcripts. NTC- non-targeting control. Relative fold change C9orf72 V3/HPRT1 1.5 1.0 0.5 0.0 0.001 0.01 0.1 1 10 Relative fold change C9orf72 V3/HPRT1 0.016 0.08 0.4 2 10 0.016 0.08 0.4 2 10 0.016 0.08 0.4 2 10 0.016 0.08 0.4 2 10 1.5 1.0 0.5 0.0 1.5 1.0 0.5 0.0 Relative fold change C9orf72/HPRT1 Neuro C9orf72
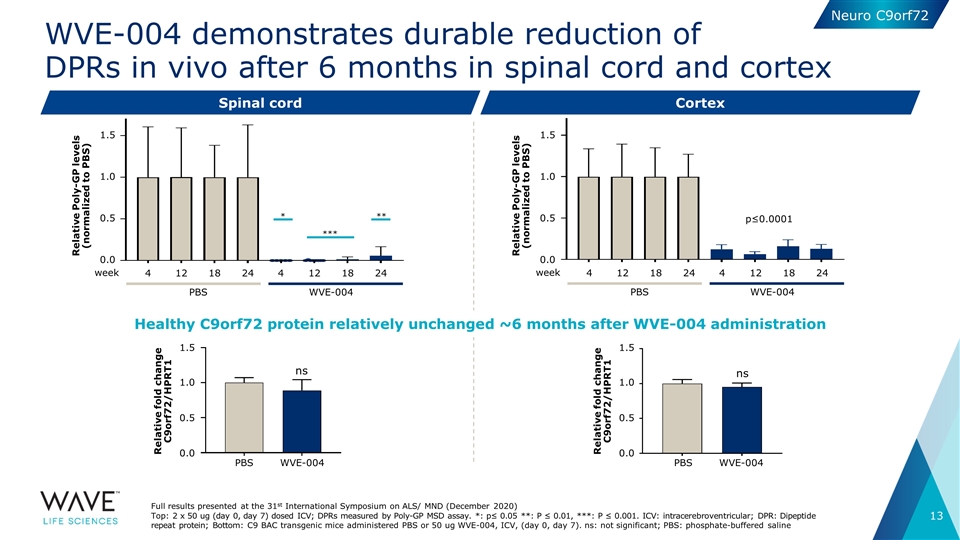
WVE-004 demonstrates durable reduction of DPRs in vivo after 6 months in spinal cord and cortex Spinal cord Cortex Full results presented at the 31st International Symposium on ALS/ MND (December 2020) Top: 2 x 50 ug (day 0, day 7) dosed ICV; DPRs measured by Poly-GP MSD assay. *: p≤ 0.05 **: P ≤ 0.01, ***: P ≤ 0.001. ICV: intracerebroventricular; DPR: Dipeptide repeat protein; Bottom: C9 BAC transgenic mice administered PBS or 50 ug WVE-004, ICV, (day 0, day 7). ns: not significant; PBS: phosphate-buffered saline * *** ** 4 12 18 12 18 24 4 24 WVE-004 PBS week 1.5 0.5 0.0 1.0 Relative Poly-GP levels (normalized to PBS) p≤0.0001 4 12 18 12 18 24 4 24 WVE-004 PBS week 1.5 0.5 0.0 1.0 Relative Poly-GP levels (normalized to PBS) ns Relative fold change C9orf72/HPRT1 1.5 0.5 0.0 1.0 WVE-004 PBS ns Relative fold change C9orf72/HPRT1 1.5 0.5 0.0 1.0 WVE-004 PBS Healthy C9orf72 protein relatively unchanged ~6 months after WVE-004 administration Neuro C9orf72
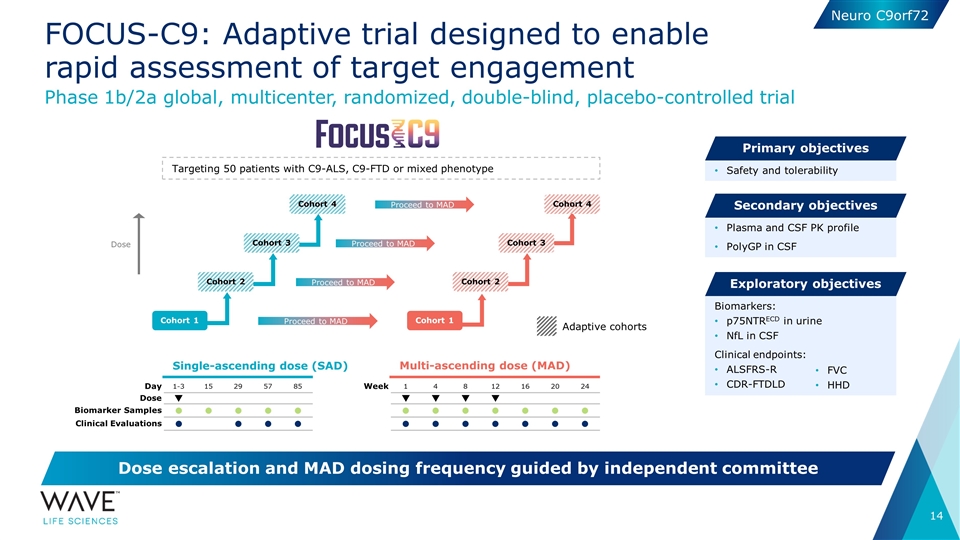
Single-ascending dose (SAD) Multi-ascending dose (MAD) Phase 1b/2a global, multicenter, randomized, double-blind, placebo-controlled trial Dose escalation and MAD dosing frequency guided by independent committee Safety and tolerability Primary objectives Plasma and CSF PK profile PolyGP in CSF Secondary objectives Biomarkers: p75NTRECD in urine NfL in CSF Clinical endpoints: ALSFRS-R CDR-FTDLD Exploratory objectives Day 1-3 15 29 57 85 Dose q Biomarker Samples l l l l l Clinical Evaluations l l l l Week 1 4 8 12 16 20 24 q q q q l l l l l l l l l l l l l l FOCUS-C9: Adaptive trial designed to enable rapid assessment of target engagement Targeting 50 patients with C9-ALS, C9-FTD or mixed phenotype Cohort 1 Proceed to MAD Proceed to MAD Proceed to MAD Cohort 4 Proceed to MAD FVC HHD Adaptive cohorts Cohort 2 Cohort 3 Cohort 1 Cohort 2 Cohort 3 Cohort 4 Dose Neuro C9orf72

WVE-003 Huntington’s Disease
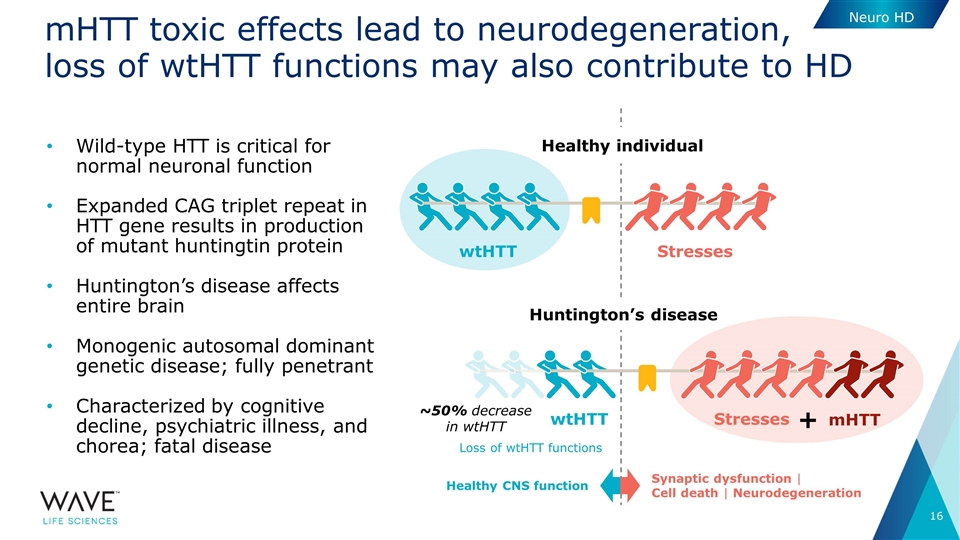
Healthy individual Huntington’s disease mHTT toxic effects lead to neurodegeneration, loss of wtHTT functions may also contribute to HD Wild-type HTT is critical for normal neuronal function Expanded CAG triplet repeat in HTT gene results in production of mutant huntingtin protein Huntington’s disease affects entire brain Monogenic autosomal dominant genetic disease; fully penetrant Characterized by cognitive decline, psychiatric illness, and chorea; fatal disease Stresses wtHTT Stresses wtHTT mHTT + ~50% decrease in wtHTT Healthy CNS function Synaptic dysfunction | Cell death | Neurodegeneration Loss of wtHTT functions Neuro HD
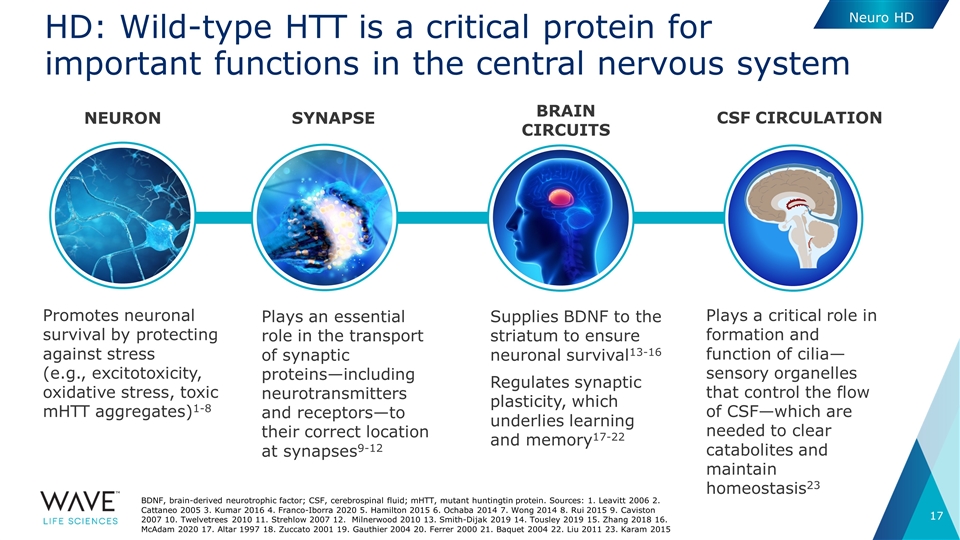
Plays an essential role in the transport of synaptic proteins—including neurotransmitters and receptors—to their correct location at synapses9-12 Promotes neuronal survival by protecting against stress (e.g., excitotoxicity, oxidative stress, toxic mHTT aggregates)1-8 BRAIN CIRCUITS SYNAPSE NEURON CSF circulation Supplies BDNF to the striatum to ensure neuronal survival13-16 Regulates synaptic plasticity, which underlies learning and memory17-22 Plays a critical role in formation and function of cilia—sensory organelles that control the flow of CSF—which are needed to clear catabolites and maintain homeostasis23 HD: Wild-type HTT is a critical protein for important functions in the central nervous system BDNF, brain-derived neurotrophic factor; CSF, cerebrospinal fluid; mHTT, mutant huntingtin protein. Sources: 1. Leavitt 2006 2. Cattaneo 2005 3. Kumar 2016 4. Franco-Iborra 2020 5. Hamilton 2015 6. Ochaba 2014 7. Wong 2014 8. Rui 2015 9. Caviston 2007 10. Twelvetrees 2010 11. Strehlow 2007 12. Milnerwood 2010 13. Smith-Dijak 2019 14. Tousley 2019 15. Zhang 2018 16. McAdam 2020 17. Altar 1997 18. Zuccato 2001 19. Gauthier 2004 20. Ferrer 2000 21. Baquet 2004 22. Liu 2011 23. Karam 2015 Neuro HD

Cerebral cortex Striatum BDNF- containing vesicle To the striatum Microtubule HTT HTT provides BDNF, a growth factor critical for survival of striatal neurons BDNF, brain-derived neurotrophic factor; HD, Huntington’s disease; HTT, huntingtin protein. 1. Altar CA, Cai N, Bliven T, et al. Nature. 1997;389(6653):856-860. 2. Zuccato C, Ciammola A, Rigamonti D, et al. Science. 2001;293(5529):493-498. 3. Gauthier LR, Charrin BC, Borrell-Pagès M, et al. Cell. 2004;118(1):127-138. 4. Ferrer I, Goutan E, Marín C, et al. Brain Res. 2000;866(1-2):257-261. 5. Baquet ZC, Gorski JA, Jones KR. J Neurosci. 2004;24(17):4250-4258. 6. Cattaneo E, et al. Nat Rev Neurosci. 2005;6(12):919-930. From the cerebral cortex Striatal neurons do not produce BDNF, but they need it to survive1 HTT promotes the production of BDNF and transports BDNF from the cortex to the striatum2,3 In HD, decreased levels of BDNF contribute to degeneration of corticostriatal circuits2,4,5 Reduction of wtHTT may decrease the availability of BDNF and accelerate corticostriatal degeneration6 Corticostriatal circuits Neuro HD
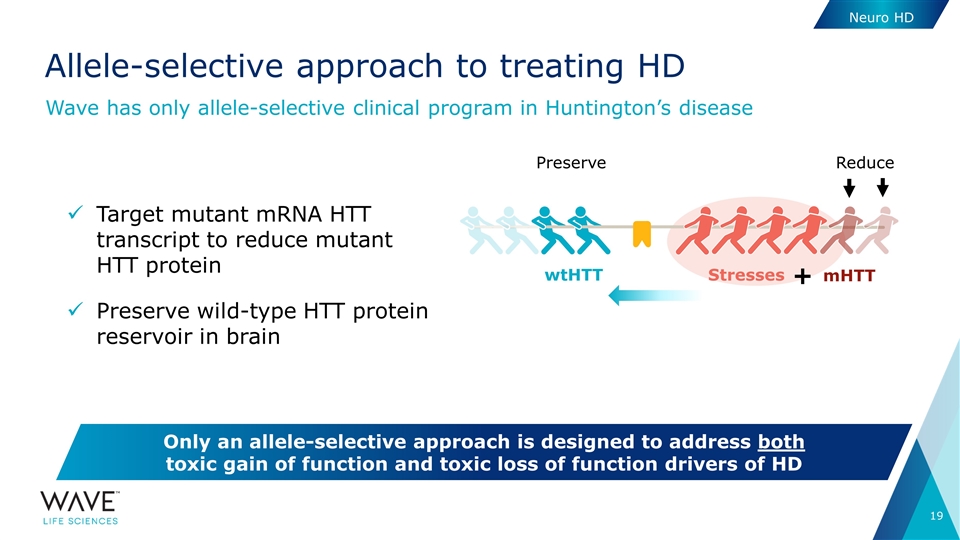
Target mutant mRNA HTT transcript to reduce mutant HTT protein Preserve wild-type HTT protein reservoir in brain Allele-selective approach to treating HD Wave has only allele-selective clinical program in Huntington’s disease Only an allele-selective approach is designed to address both toxic gain of function and toxic loss of function drivers of HD Stresses wtHTT mHTT + Reduce Preserve Neuro HD
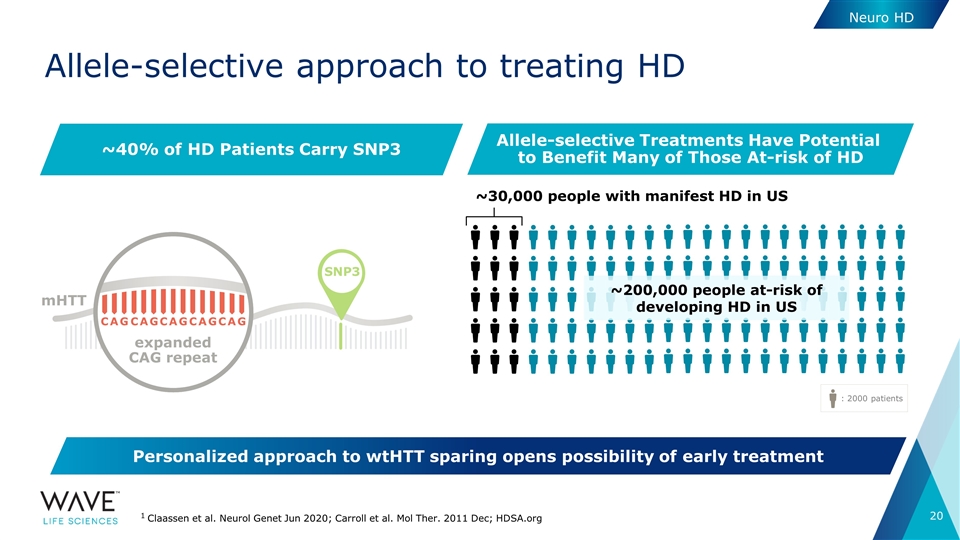
Allele-selective approach to treating HD Neuro HD 1 Claassen et al. Neurol Genet Jun 2020; Carroll et al. Mol Ther. 2011 Dec; HDSA.org : 2000 patients ~30,000 people with manifest HD in US ~40% of HD Patients Carry SNP3 Allele-selective Treatments Have Potential to Benefit Many of Those At-risk of HD ~200,000 people at-risk of developing HD in US SNP3 C A G C A G C A G C A G C A G expanded CAG repeat mHTT Personalized approach to wtHTT sparing opens possibility of early treatment
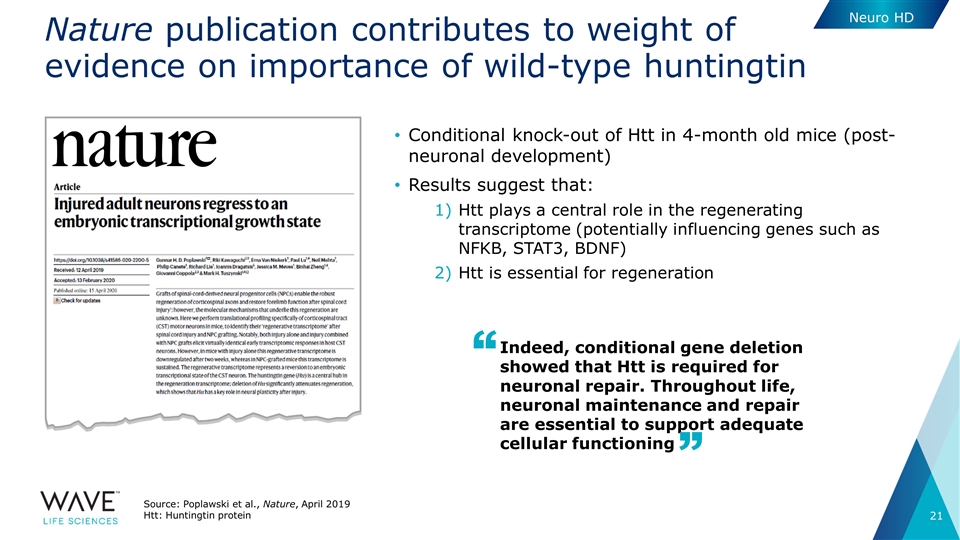

Nature publication contributes to weight of evidence on importance of wild-type huntingtin Source: Poplawski et al., Nature, April 2019 Htt: Huntingtin protein Conditional knock-out of Htt in 4-month old mice (post-neuronal development) Results suggest that: Htt plays a central role in the regenerating transcriptome (potentially influencing genes such as NFKB, STAT3, BDNF) Htt is essential for regeneration Indeed, conditional gene deletion showed that Htt is required for neuronal repair. Throughout life, neuronal maintenance and repair are essential to support adequate cellular functioning Neuro HD
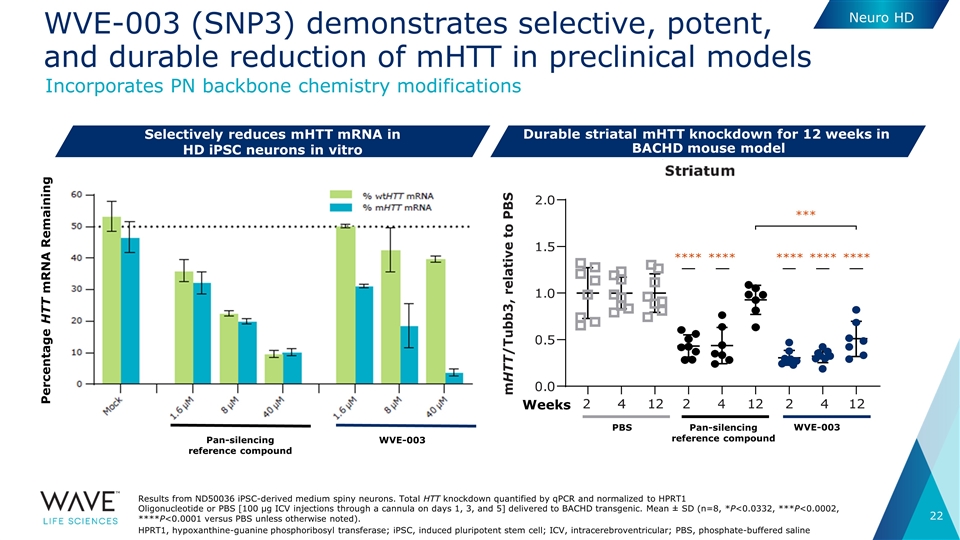
WVE-003 (SNP3) demonstrates selective, potent, and durable reduction of mHTT in preclinical models Selectively reduces mHTT mRNA in HD iPSC neurons in vitro Results from ND50036 iPSC-derived medium spiny neurons. Total HTT knockdown quantified by qPCR and normalized to HPRT1 Oligonucleotide or PBS [100 μg ICV injections through a cannula on days 1, 3, and 5] delivered to BACHD transgenic. Mean ± SD (n=8, *P<0.0332, ***P<0.0002, ****P<0.0001 versus PBS unless otherwise noted). HPRT1, hypoxanthine-guanine phosphoribosyl transferase; iPSC, induced pluripotent stem cell; ICV, intracerebroventricular; PBS, phosphate-buffered saline Similar results in cortex Pan-silencing reference compound WVE-003 PBS Weeks *** **** **** **** **** **** Pan-silencing reference compound WVE-003 Percentage HTT mRNA Remaining Durable striatal mHTT knockdown for 12 weeks in BACHD mouse model Neuro HD Incorporates PN backbone chemistry modifications
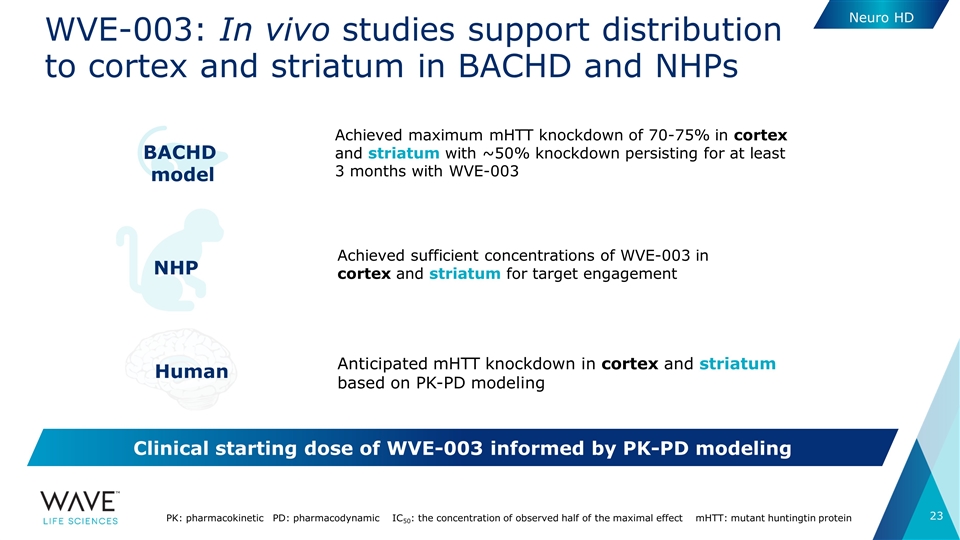

WVE-003: In vivo studies support distribution to cortex and striatum in BACHD and NHPs PK: pharmacokinetic PD: pharmacodynamic IC50: the concentration of observed half of the maximal effect mHTT: mutant huntingtin protein Achieved sufficient concentrations of WVE-003 in cortex and striatum for target engagement NHP Anticipated mHTT knockdown in cortex and striatum based on PK-PD modeling Human BACHD model Achieved maximum mHTT knockdown of 70-75% in cortex and striatum with ~50% knockdown persisting for at least 3 months with WVE-003 Clinical starting dose of WVE-003 informed by PK-PD modeling Neuro HD
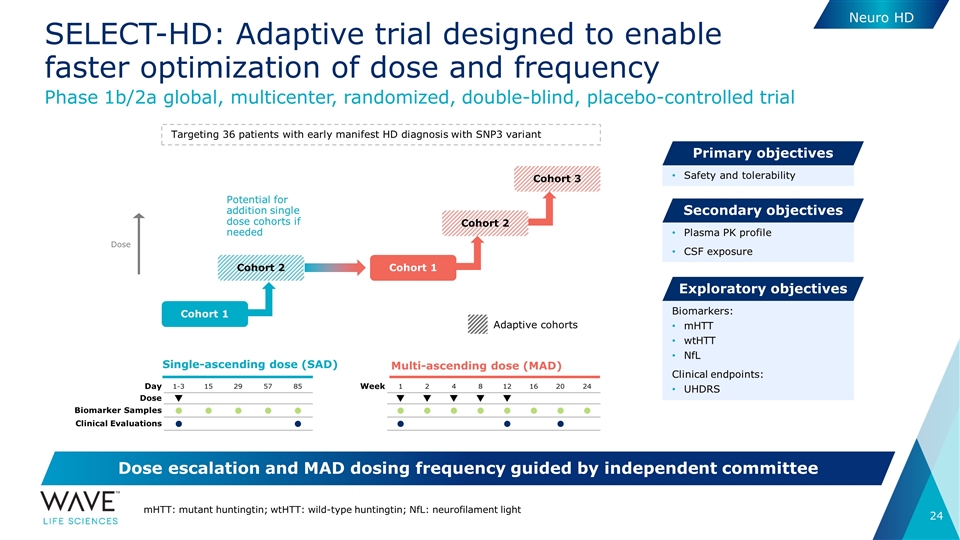
Single-ascending dose (SAD) Multi-ascending dose (MAD) Day 1-3 15 29 57 85 Dose q Biomarker Samples l l l l l Clinical Evaluations l l Week 1 2 4 8 12 16 20 24 q q q q q l l l l l l l l l l l SELECT-HD: Adaptive trial designed to enable faster optimization of dose and frequency Adaptive cohorts Phase 1b/2a global, multicenter, randomized, double-blind, placebo-controlled trial Safety and tolerability Primary objectives Plasma PK profile CSF exposure Secondary objectives Biomarkers: mHTT wtHTT NfL Clinical endpoints: UHDRS Exploratory objectives Dose escalation and MAD dosing frequency guided by independent committee mHTT: mutant huntingtin; wtHTT: wild-type huntingtin; NfL: neurofilament light Targeting 36 patients with early manifest HD diagnosis with SNP3 variant Cohort 1 Cohort 2 Cohort 1 Cohort 2 Cohort 3 Potential for addition single dose cohorts if needed Dose Neuro HD
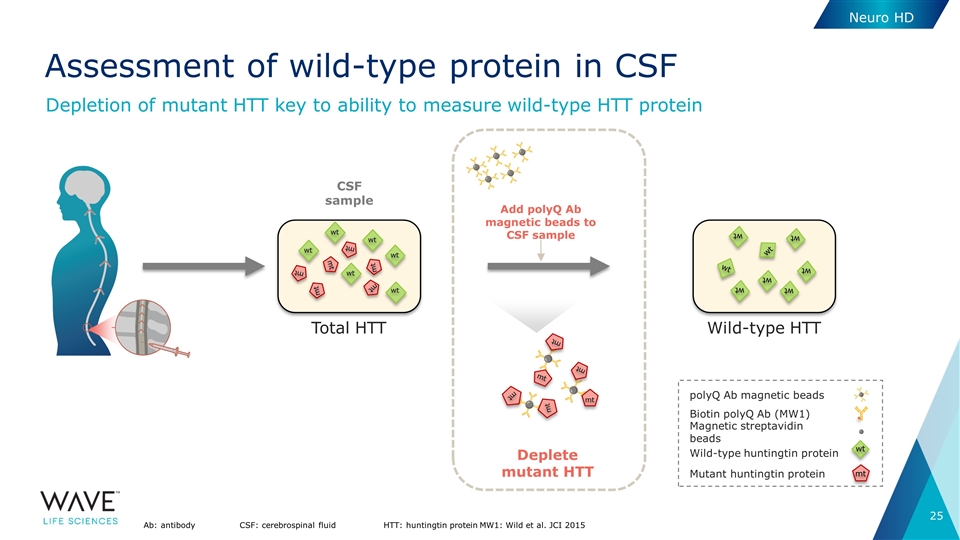
Neuro HD Assessment of wild-type protein in CSF Ab: antibodyCSF: cerebrospinal fluidHTT: huntingtin proteinMW1: Wild et al. JCI 2015 CSF sample Total HTT mt wt wt wt mt mt mt mt mt wt wt wt Wild-type HTT wt wt wt wt wt wt wt wt Deplete mutant HTT Add polyQ Ab magnetic beads to CSF sample mt mt mt mt mt mt polyQ Ab magnetic beads Biotin polyQ Ab (MW1) Magnetic streptavidin beads Wild-type huntingtin protein Mutant huntingtin protein wt mt Depletion of mutant HTT key to ability to measure wild-type HTT protein

WVE-N531 Duchenne muscular dystrophy
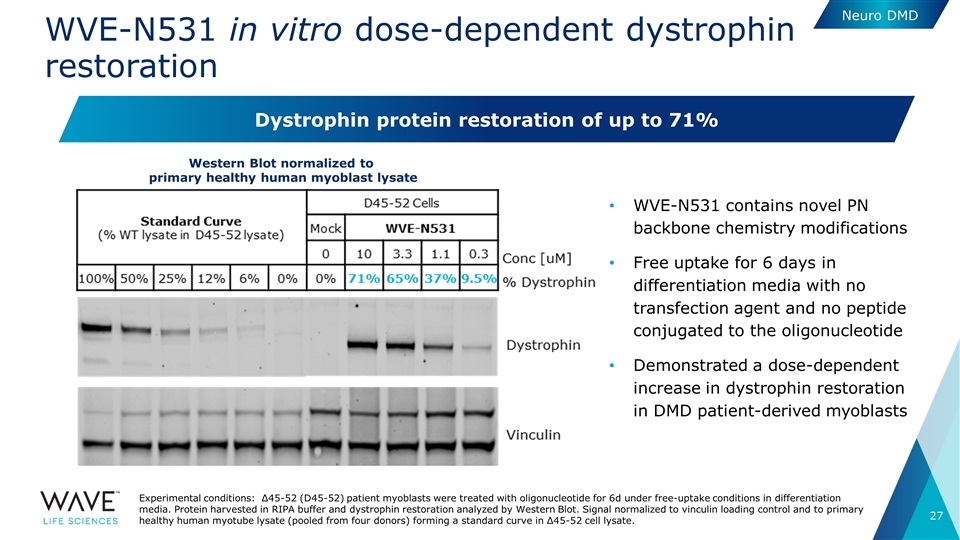
WVE-N531 in vitro dose-dependent dystrophin restoration WVE-N531 contains novel PN backbone chemistry modifications Free uptake for 6 days in differentiation media with no transfection agent and no peptide conjugated to the oligonucleotide Demonstrated a dose-dependent increase in dystrophin restoration in DMD patient-derived myoblasts Experimental conditions: Δ45-52 (D45-52) patient myoblasts were treated with oligonucleotide for 6d under free-uptake conditions in differentiation media. Protein harvested in RIPA buffer and dystrophin restoration analyzed by Western Blot. Signal normalized to vinculin loading control and to primary healthy human myotube lysate (pooled from four donors) forming a standard curve in Δ45-52 cell lysate. Western Blot normalized to primary healthy human myoblast lysate Dystrophin protein restoration of up to 71% Neuro DMD
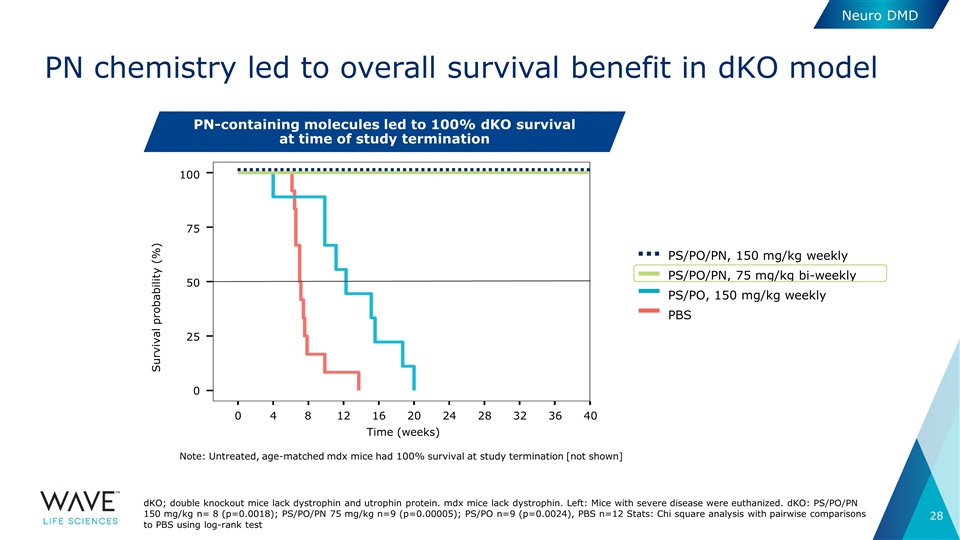
PN chemistry led to overall survival benefit in dKO model dKO; double knockout mice lack dystrophin and utrophin protein. mdx mice lack dystrophin. Left: Mice with severe disease were euthanized. dKO: PS/PO/PN 150 mg/kg n= 8 (p=0.0018); PS/PO/PN 75 mg/kg n=9 (p=0.00005); PS/PO n=9 (p=0.0024), PBS n=12 Stats: Chi square analysis with pairwise comparisons to PBS using log-rank test PN-containing molecules led to 100% dKO survival at time of study termination PS/PO/PN, 75 mg/kg bi-weekly PBS PS/PO, 150 mg/kg weekly PS/PO/PN, 150 mg/kg weekly 100 75 50 25 0 Survival probability (%) 0 4 8 12 16 20 24 28 32 36 40 Time (weeks) Note: Untreated, age-matched mdx mice had 100% survival at study termination [not shown] Neuro DMD

Clinical trial of WVE-N531 to initiate in 2021 Unmet need in DMD remains high CTA submitted in March 2021 to initiate clinical development Clinical trial powered to evaluate change in dystrophin production, and will assess drug concentration in muscle, and initial safety Open-label study; targeting every-other-week administration in up to 15 boys with DMD Potential to apply PN chemistry to other exons if successful Dosing in clinical trial expected to initiate in 2021 Neuro DMD

Wave’s discovery and drug development platform
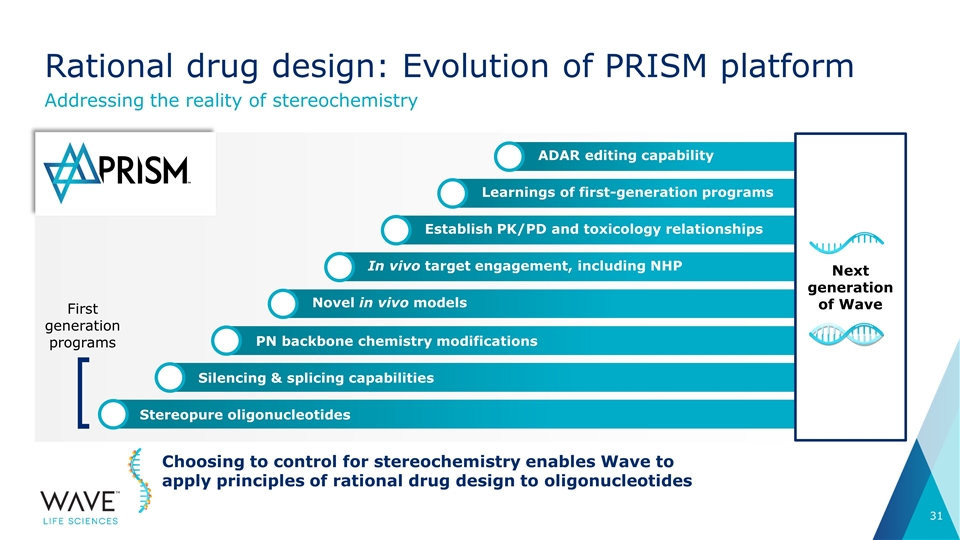
Silencing & splicing capabilities PN backbone chemistry modifications In vivo target engagement, including NHP ADAR editing capability Learnings of first-generation programs Establish PK/PD and toxicology relationships Novel in vivo models Stereopure oligonucleotides Rational drug design: Evolution of PRISM platform Addressing the reality of stereochemistry Choosing to control for stereochemistry enables Wave to apply principles of rational drug design to oligonucleotides First generation programs Next generation of Wave
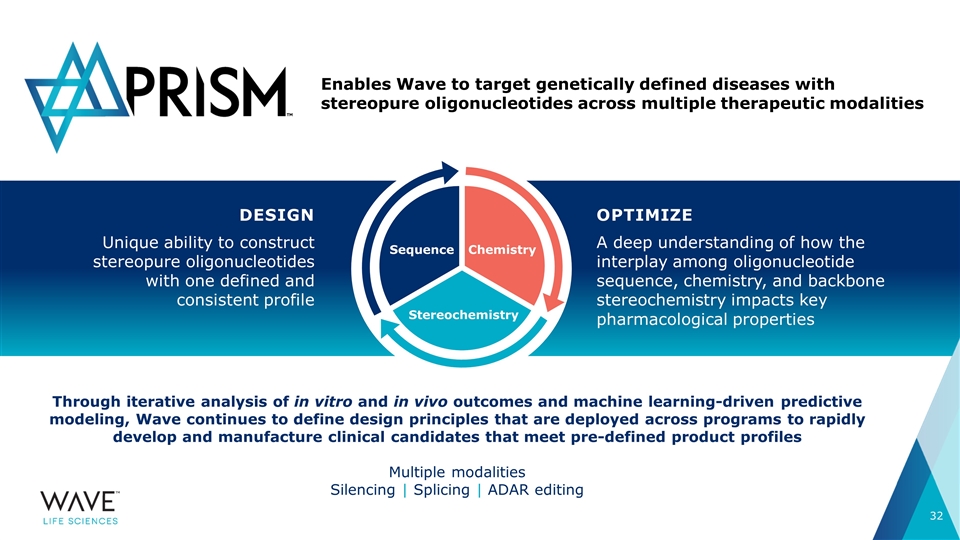
Through iterative analysis of in vitro and in vivo outcomes and machine learning-driven predictive modeling, Wave continues to define design principles that are deployed across programs to rapidly develop and manufacture clinical candidates that meet pre-defined product profiles Multiple modalities Silencing | Splicing | ADAR editing DESIGN Unique ability to construct stereopure oligonucleotides with one defined and consistent profile Enables Wave to target genetically defined diseases with stereopure oligonucleotides across multiple therapeutic modalities OPTIMIZE A deep understanding of how the interplay among oligonucleotide sequence, chemistry, and backbone stereochemistry impacts key pharmacological properties SEQUENCE STEREOCHEMISTRY CHEMISTRY Sequence Stereochemistry Chemistry
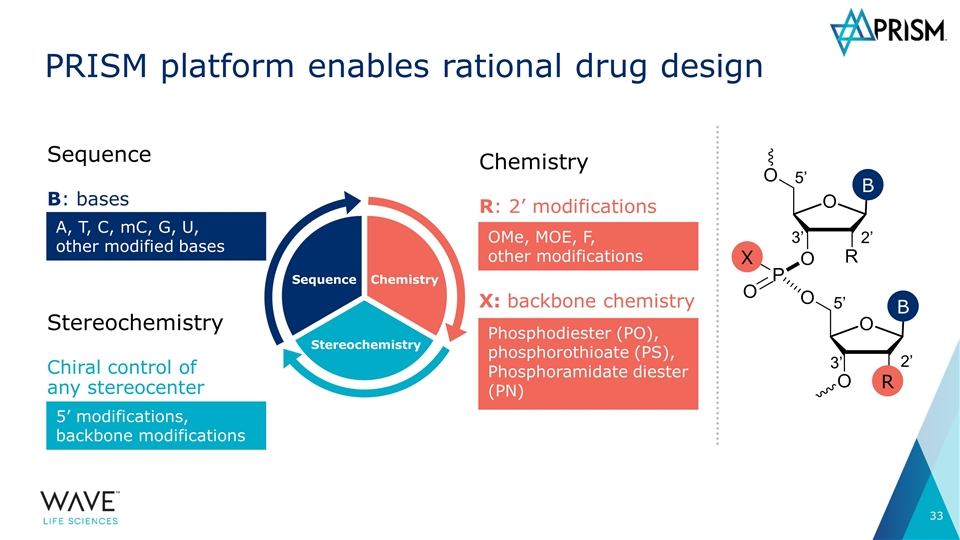
Sequence Stereochemistry Chemistry PRISM platform enables rational drug design Chemistry R: 2’ modifications OMe, MOE, F, other modifications 5’ 2’ 3’ 5’ 3’ 2’ R X B B X: backbone chemistry Phosphodiester (PO), phosphorothioate (PS), Phosphoramidate diester (PN) Sequence B: bases A, T, C, mC, G, U, other modified bases Stereochemistry Chiral control of any stereocenter 5’ modifications, backbone modifications
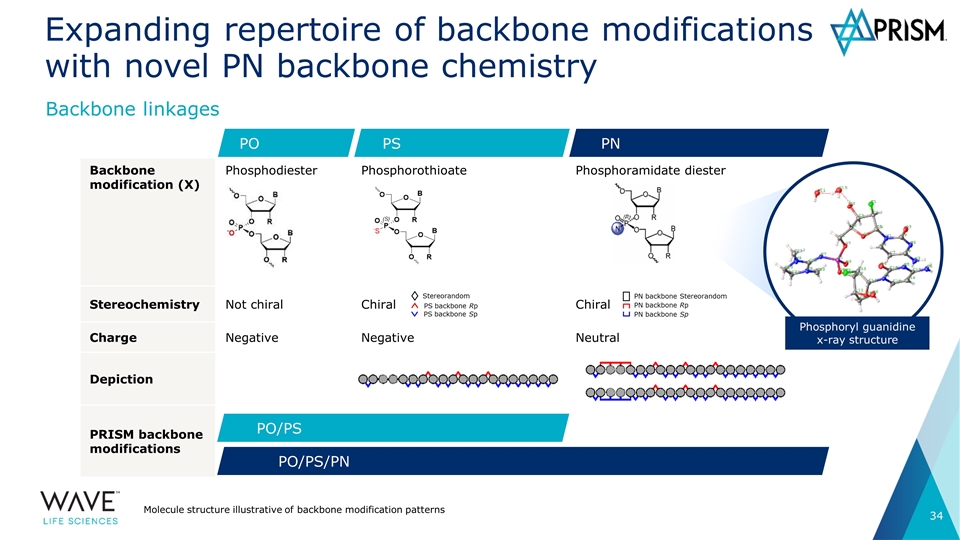
Backbone modification (X) Phosphodiester Phosphorothioate Phosphoramidate diester Stereochemistry Not chiral Chiral Chiral Charge Negative Negative Neutral Depiction PRISM backbone modifications Expanding repertoire of backbone modifications with novel PN backbone chemistry Molecule structure illustrative of backbone modification patterns Backbone linkages PS PO PN PO/PS PO/PS/PN Phosphoryl guanidine x-ray structure Stereorandom PS backbone Rp PS backbone Sp PN backbone Sp PN backbone Rp PN backbone Stereorandom
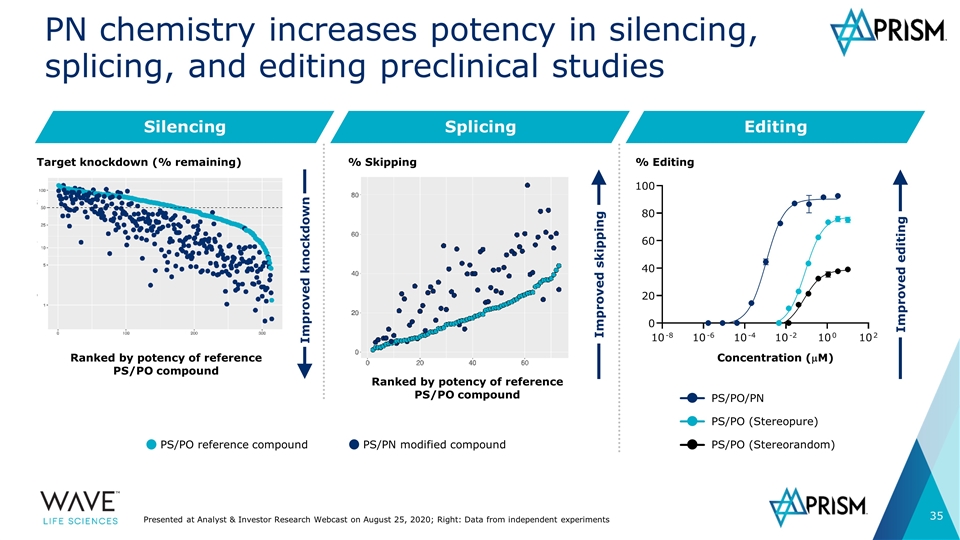
Silencing PN chemistry increases potency in silencing, splicing, and editing preclinical studies Improved knockdown Splicing Editing Improved skipping Ranked by potency of reference PS/PO compound Ranked by potency of reference PS/PO compound Improved editing PS/PO/PN PS/PO (Stereopure) PS/PO (Stereorandom) Concentration (mM) % Editing PS/PO reference compound PS/PN modified compound % Skipping Target knockdown (% remaining) Presented at Analyst & Investor Research Webcast on August 25, 2020; Right: Data from independent experiments
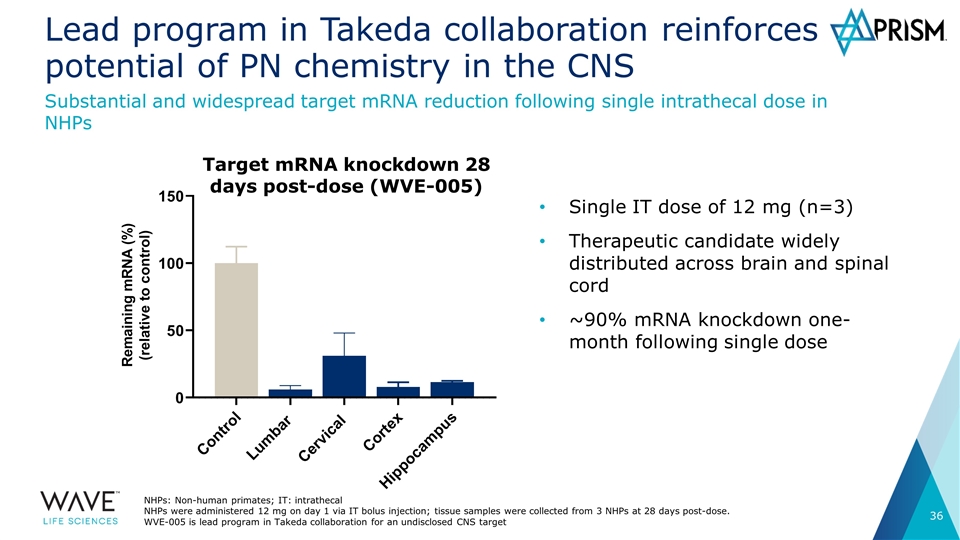
Lead program in Takeda collaboration reinforces potential of PN chemistry in the CNS Single IT dose of 12 mg (n=3) Therapeutic candidate widely distributed across brain and spinal cord ~90% mRNA knockdown one-month following single dose Substantial and widespread target mRNA reduction following single intrathecal dose in NHPs NHPs: Non-human primates; IT: intrathecal NHPs were administered 12 mg on day 1 via IT bolus injection; tissue samples were collected from 3 NHPs at 28 days post-dose. WVE-005 is lead program in Takeda collaboration for an undisclosed CNS target Target mRNA knockdown 28 days post-dose (WVE-005)
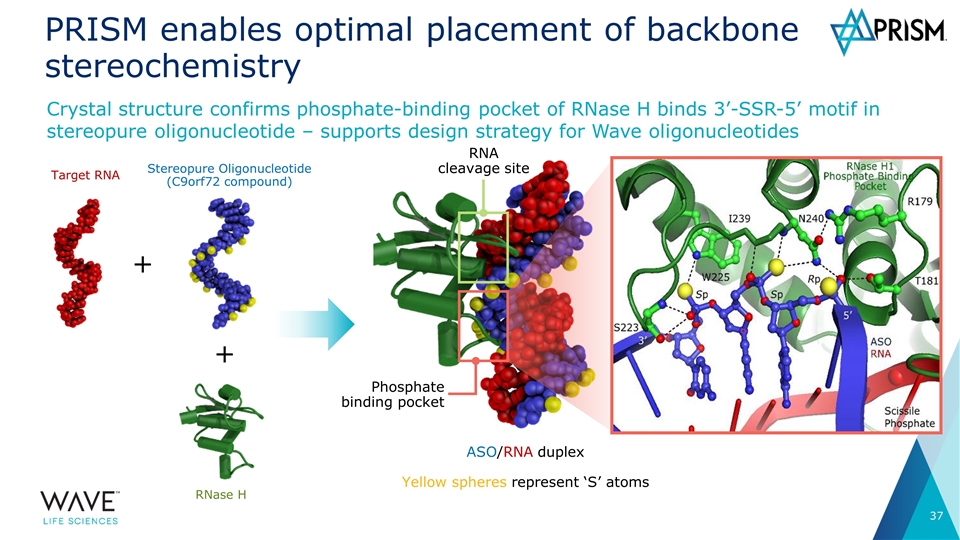
PRISM enables optimal placement of backbone stereochemistry Crystal structure confirms phosphate-binding pocket of RNase H binds 3’-SSR-5’ motif in stereopure oligonucleotide – supports design strategy for Wave oligonucleotides ASO/RNA duplex Yellow spheres represent ‘S’ atoms Phosphate binding pocket RNA cleavage site Target RNA Stereopure Oligonucleotide (C9orf72 compound) RNase H + +
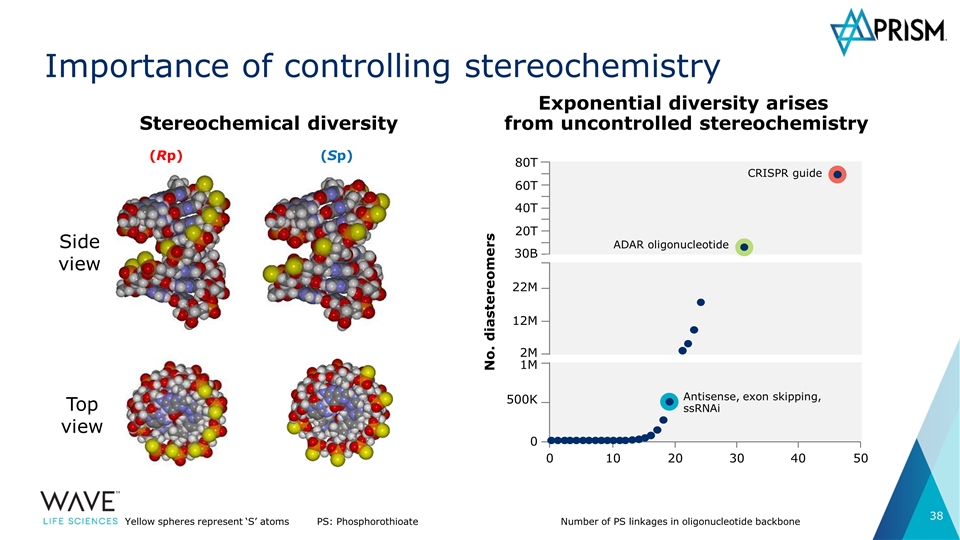
Importance of controlling stereochemistry (Rp) (Sp) Top view Side view Yellow spheres represent ‘S’ atomsPS: Phosphorothioate Number of PS linkages in oligonucleotide backbone No. diastereomers 80T 60T 40T 20T 30B 22M 12M 2M 1M 500K 0 0 10 20 30 40 50 Antisense, exon skipping, ssRNAi ADAR oligonucleotide CRISPR guide Stereochemical diversity Exponential diversity arises from uncontrolled stereochemistry

ADAR editing Platform capability and Alpha-1 antitrypsin deficiency
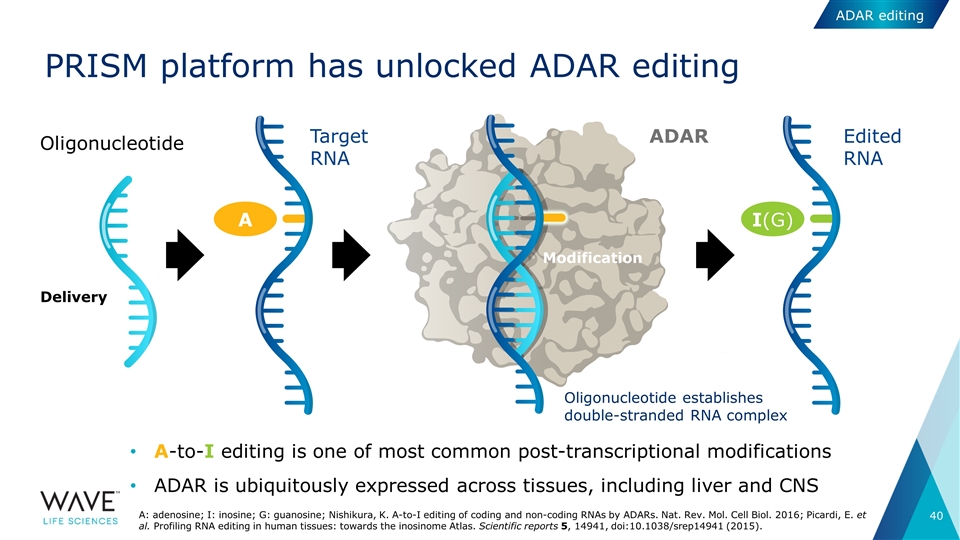
PRISM platform has unlocked ADAR editing A-to-I editing is one of most common post-transcriptional modifications ADAR is ubiquitously expressed across tissues, including liver and CNS ADAR Target RNA I(G) A Edited RNA Oligonucleotide establishes double-stranded RNA complex Oligonucleotide Modification Delivery A: adenosine; I: inosine; G: guanosine; Nishikura, K. A-to-I editing of coding and non-coding RNAs by ADARs. Nat. Rev. Mol. Cell Biol. 2016; Picardi, E. et al. Profiling RNA editing in human tissues: towards the inosinome Atlas. Scientific reports 5, 14941, doi:10.1038/srep14941 (2015). ADAR editing
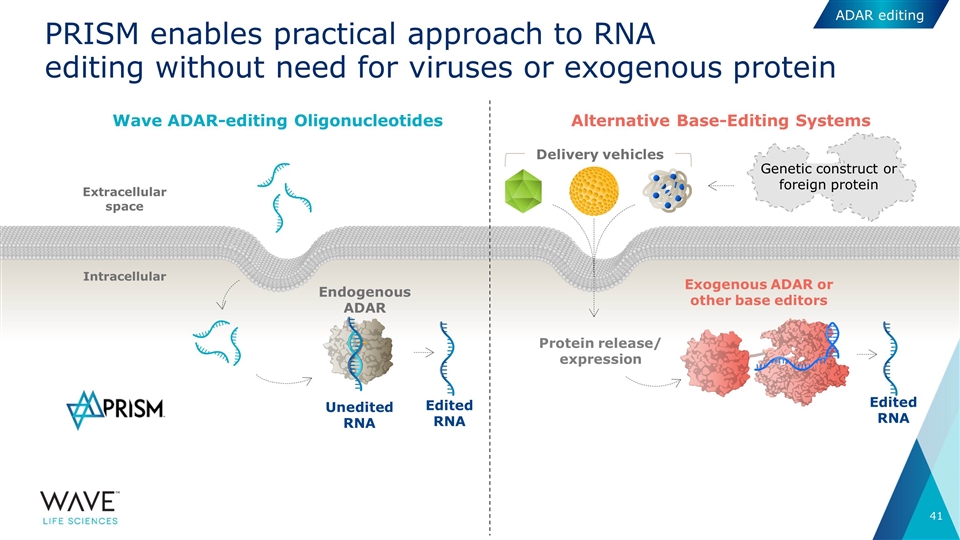
PRISM enables practical approach to RNA editing without need for viruses or exogenous protein Intracellular Extracellular space Endogenous ADAR Unedited RNA Wave ADAR-editing Oligonucleotides Exogenous ADAR or other base editors Edited RNA Protein release/ expression Delivery vehicles Alternative Base-Editing Systems Edited RNA Genetic construct or foreign protein ADAR editing
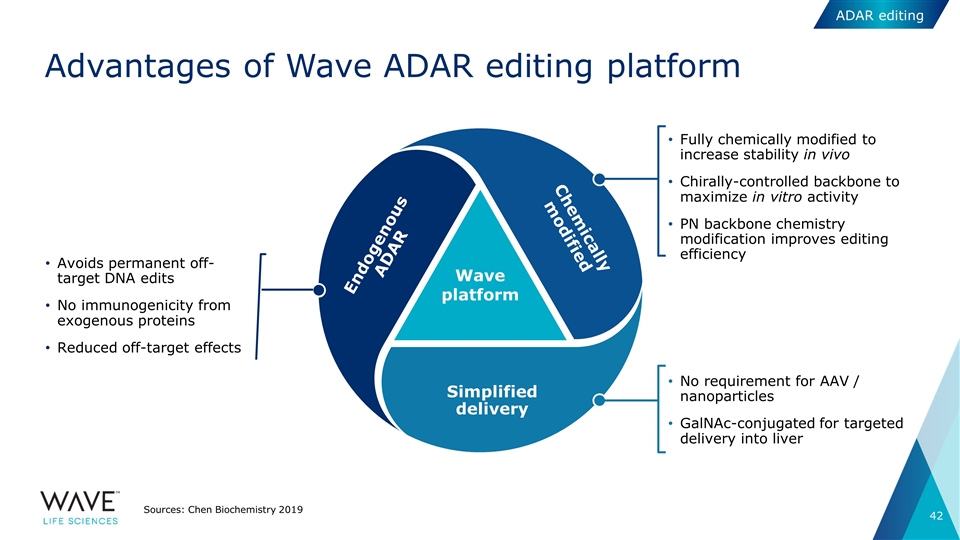
Wave platform Fully chemically modified to increase stability in vivo Chirally-controlled backbone to maximize in vitro activity PN backbone chemistry modification improves editing efficiency No requirement for AAV / nanoparticles GalNAc-conjugated for targeted delivery into liver Avoids permanent off-target DNA edits No immunogenicity from exogenous proteins Reduced off-target effects Advantages of Wave ADAR editing platform Sources: Chen Biochemistry 2019 Chemically modified Simplified delivery Endogenous ADAR ADAR editing
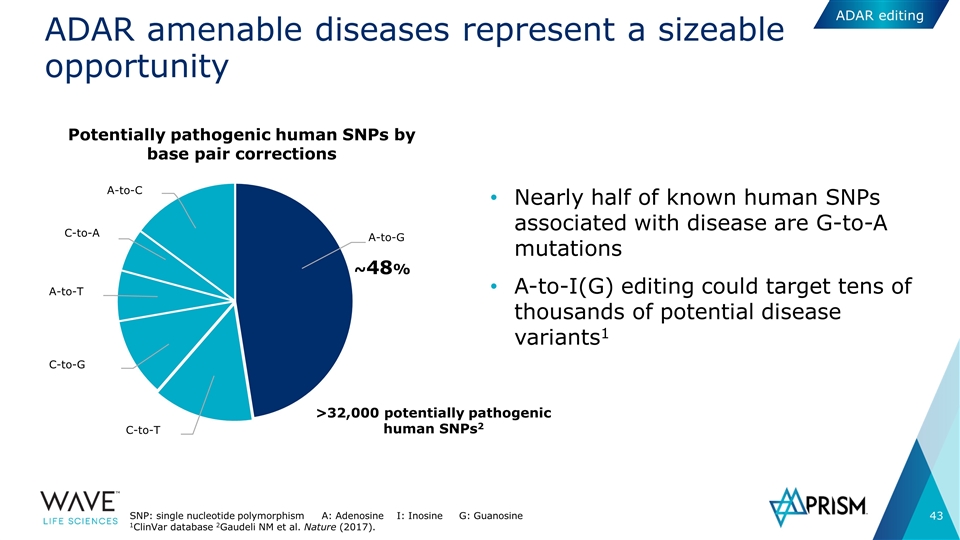
ADAR amenable diseases represent a sizeable opportunity Nearly half of known human SNPs associated with disease are G-to-A mutations A-to-I(G) editing could target tens of thousands of potential disease variants1 ~48% C-to-T C-to-G A-to-T C-to-A A-to-C A-to-G Potentially pathogenic human SNPs by base pair corrections >32,000 potentially pathogenic human SNPs2 SNP: single nucleotide polymorphism A: Adenosine I: Inosine G: Guanosine 1ClinVar database 2Gaudeli NM et al. Nature (2017). ADAR editing
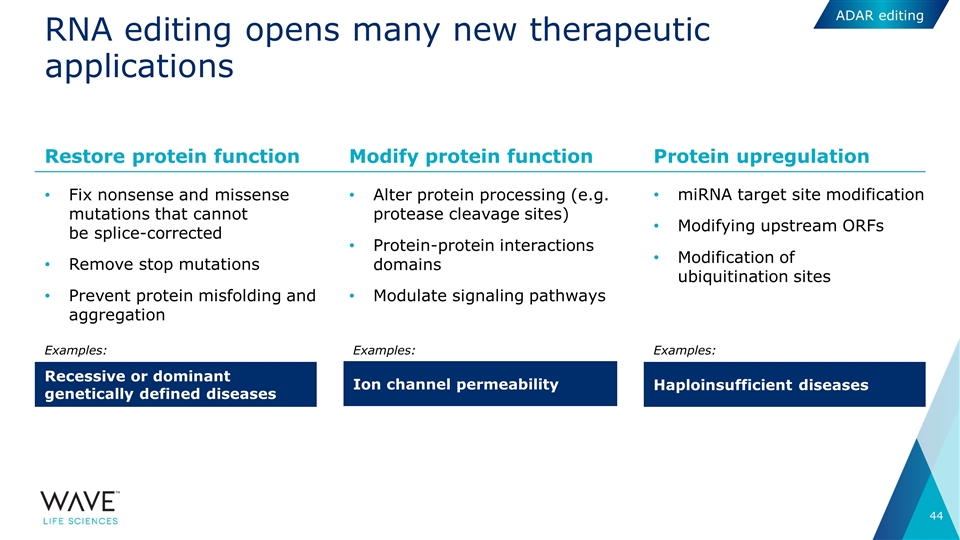
RNA editing opens many new therapeutic applications Fix nonsense and missense mutations that cannot be splice-corrected Remove stop mutations Prevent protein misfolding and aggregation Alter protein processing (e.g. protease cleavage sites) Protein-protein interactions domains Modulate signaling pathways miRNA target site modification Modifying upstream ORFs Modification of ubiquitination sites Restore protein function Recessive or dominant genetically defined diseases Modify protein function Ion channel permeability Protein upregulation Haploinsufficient diseases Examples: Examples: Examples: ADAR editing
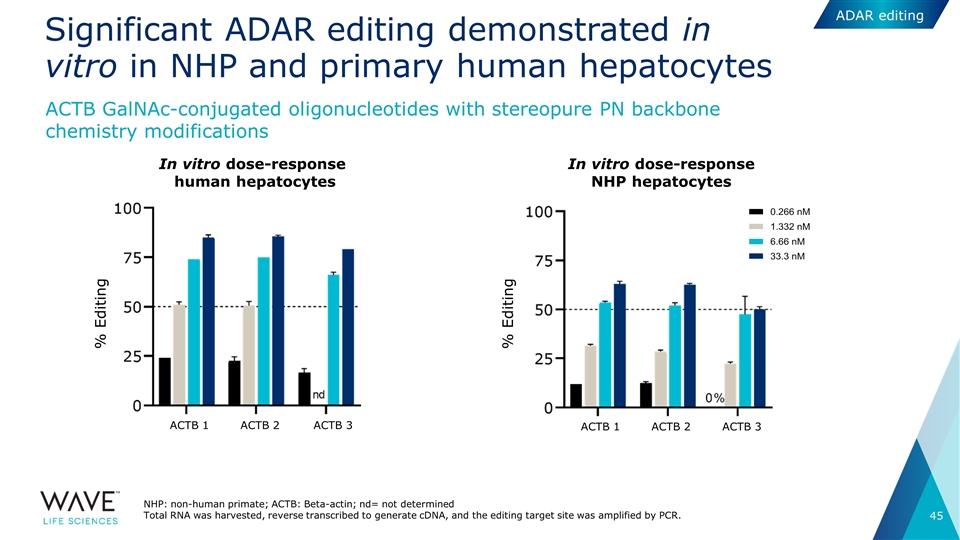
Significant ADAR editing demonstrated in vitro in NHP and primary human hepatocytes NHP: non-human primate; ACTB: Beta-actin; nd= not determined Total RNA was harvested, reverse transcribed to generate cDNA, and the editing target site was amplified by PCR. In vitro dose-response human hepatocytes In vitro dose-response NHP hepatocytes % Editing % Editing ACTB 1 ACTB 2 ACTB 3 ACTB 1 ACTB 2 ACTB 3 ACTB GalNAc-conjugated oligonucleotides with stereopure PN backbone chemistry modifications ADAR editing
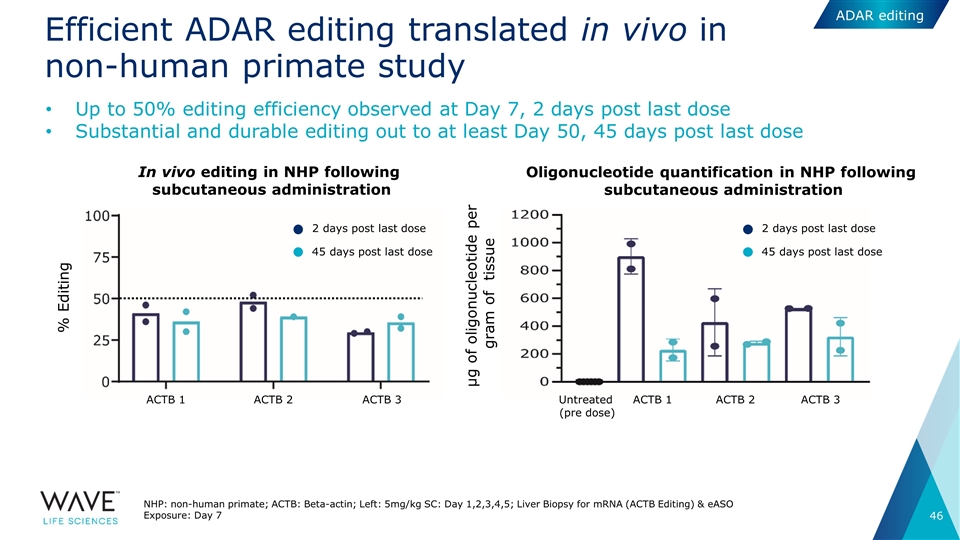

Efficient ADAR editing translated in vivo in non-human primate study NHP: non-human primate; ACTB: Beta-actin; Left: 5mg/kg SC: Day 1,2,3,4,5; Liver Biopsy for mRNA (ACTB Editing) & eASO Exposure: Day 7 Up to 50% editing efficiency observed at Day 7, 2 days post last dose Substantial and durable editing out to at least Day 50, 45 days post last dose In vivo editing in NHP following subcutaneous administration Oligonucleotide quantification in NHP following subcutaneous administration 2 days post last dose 45 days post last dose 2 days post last dose 45 days post last dose ACTB 1 ACTB 2 ACTB 3 ACTB 1 ACTB 2 ACTB 3 Untreated (pre dose) % Editing µg of oligonucleotide per gram of tissue ADAR editing
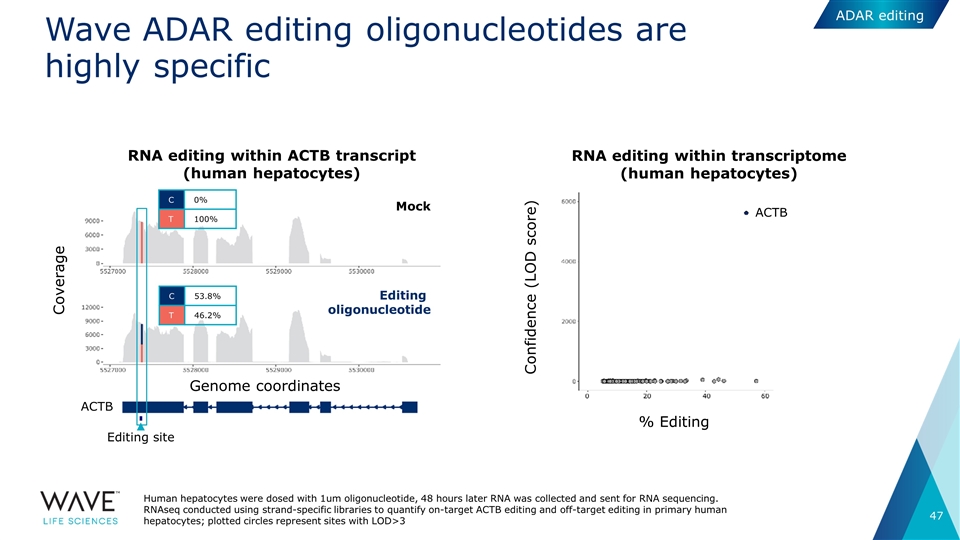

Editing site RNA editing within ACTB transcript (human hepatocytes) RNA editing within transcriptome (human hepatocytes) Wave ADAR editing oligonucleotides are highly specific Coverage Genome coordinates ACTB C 0% T 100% C 53.8% T 46.2% ACTB Confidence (LOD score) % Editing Mock Editing oligonucleotide Human hepatocytes were dosed with 1um oligonucleotide, 48 hours later RNA was collected and sent for RNA sequencing. RNAseq conducted using strand-specific libraries to quantify on-target ACTB editing and off-target editing in primary human hepatocytes; plotted circles represent sites with LOD>3 ADAR editing
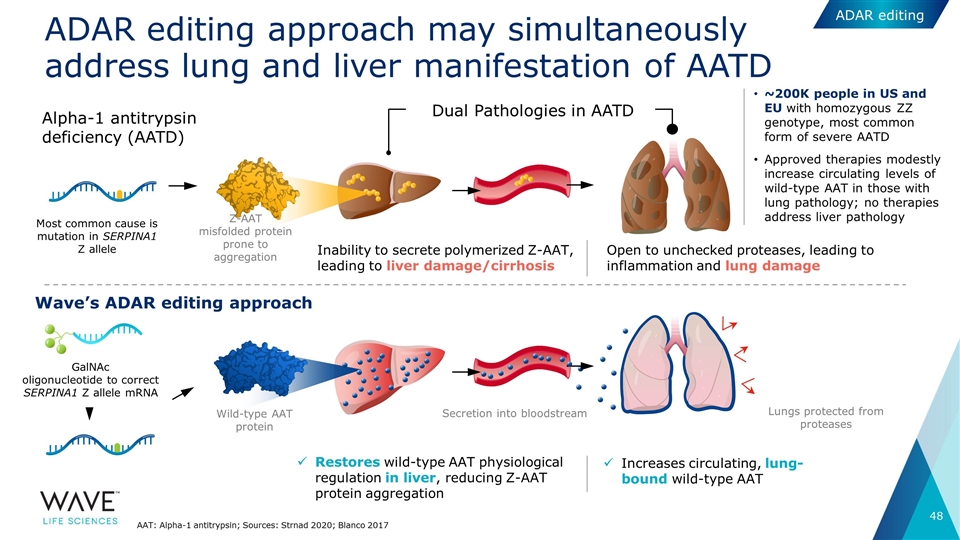
ADAR editing approach may simultaneously address lung and liver manifestation of AATD GalNAc oligonucleotide to correct SERPINA1 Z allele mRNA Most common cause is mutation in SERPINA1 Z allele Z-AAT misfolded protein prone to aggregation Inability to secrete polymerized Z-AAT, leading to liver damage/cirrhosis Open to unchecked proteases, leading to inflammation and lung damage Increases circulating, lung-bound wild-type AAT Wild-type AAT protein Secretion into bloodstream Lungs protected from proteases Restores wild-type AAT physiological regulation in liver, reducing Z-AAT protein aggregation Wave’s ADAR editing approach Alpha-1 antitrypsin deficiency (AATD) Dual Pathologies in AATD AAT: Alpha-1 antitrypsin; Sources: Strnad 2020; Blanco 2017 ~200K people in US and EU with homozygous ZZ genotype, most common form of severe AATD Approved therapies modestly increase circulating levels of wild-type AAT in those with lung pathology; no therapies address liver pathology ADAR editing
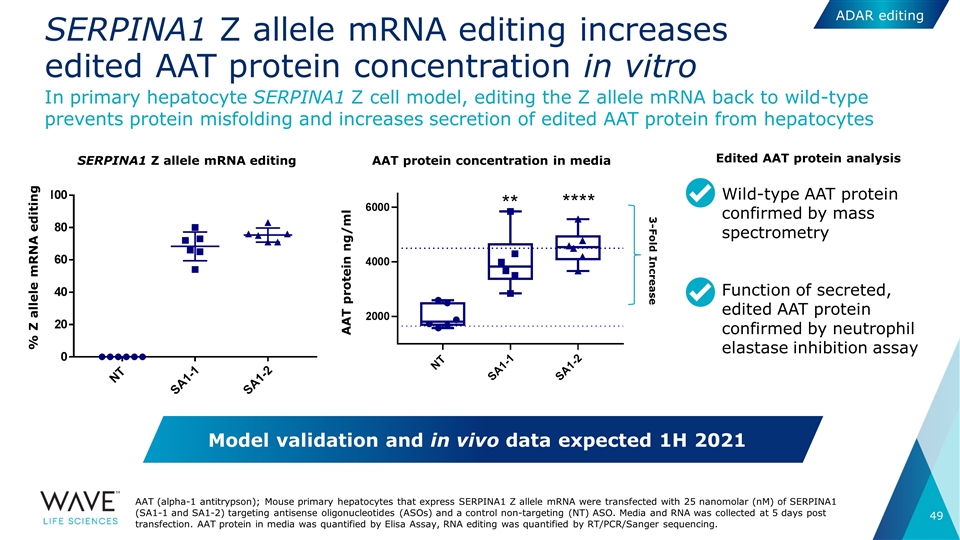

SERPINA1 Z allele mRNA editing increases edited AAT protein concentration in vitro AAT (alpha-1 antitrypson); Mouse primary hepatocytes that express SERPINA1 Z allele mRNA were transfected with 25 nanomolar (nM) of SERPINA1 (SA1-1 and SA1-2) targeting antisense oligonucleotides (ASOs) and a control non-targeting (NT) ASO. Media and RNA was collected at 5 days post transfection. AAT protein in media was quantified by Elisa Assay, RNA editing was quantified by RT/PCR/Sanger sequencing. In primary hepatocyte SERPINA1 Z cell model, editing the Z allele mRNA back to wild-type prevents protein misfolding and increases secretion of edited AAT protein from hepatocytes 3-Fold Increase AAT protein concentration in media SERPINA1 Z allele mRNA editing % Z allele mRNA editing AAT protein ng/ml Wild-type AAT protein confirmed by mass spectrometry Function of secreted, edited AAT protein confirmed by neutrophil elastase inhibition assay Model validation and in vivo data expected 1H 2021 Edited AAT protein analysis ADAR editing
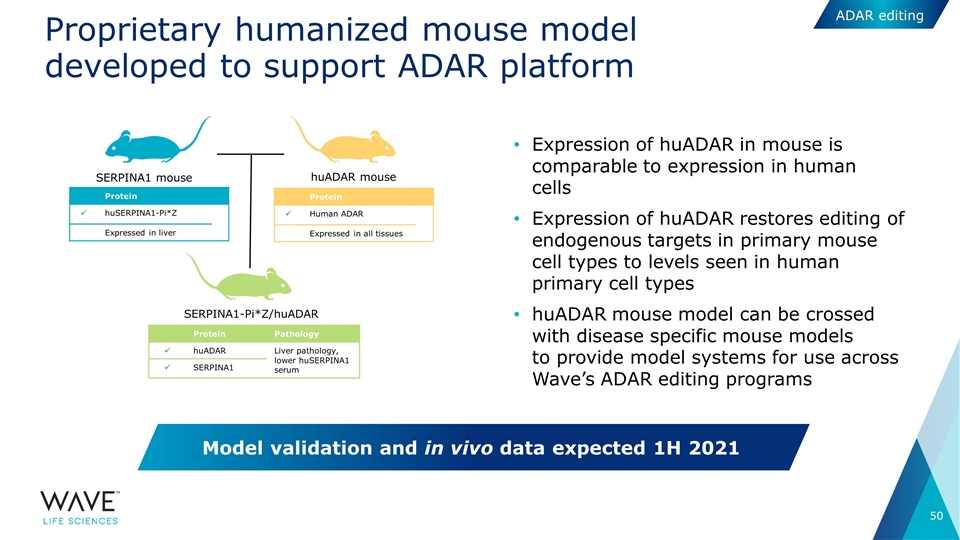
Proprietary humanized mouse model developed to support ADAR platform Model validation and in vivo data expected 1H 2021 SERPINA1-Pi*Z/huADAR Protein ü Human ADAR Expressed in all tissues huADAR mouse Protein Pathology ü huADAR Liver pathology, lower huSERPINA1 serum ü SERPINA1 SERPINA1 mouse Protein ü huSERPINA1-Pi*Z Expressed in liver Expression of huADAR in mouse is comparable to expression in human cells Expression of huADAR restores editing of endogenous targets in primary mouse cell types to levels seen in human primary cell types huADAR mouse model can be crossed with disease specific mouse models to provide model systems for use across Wave’s ADAR editing programs ADAR editing
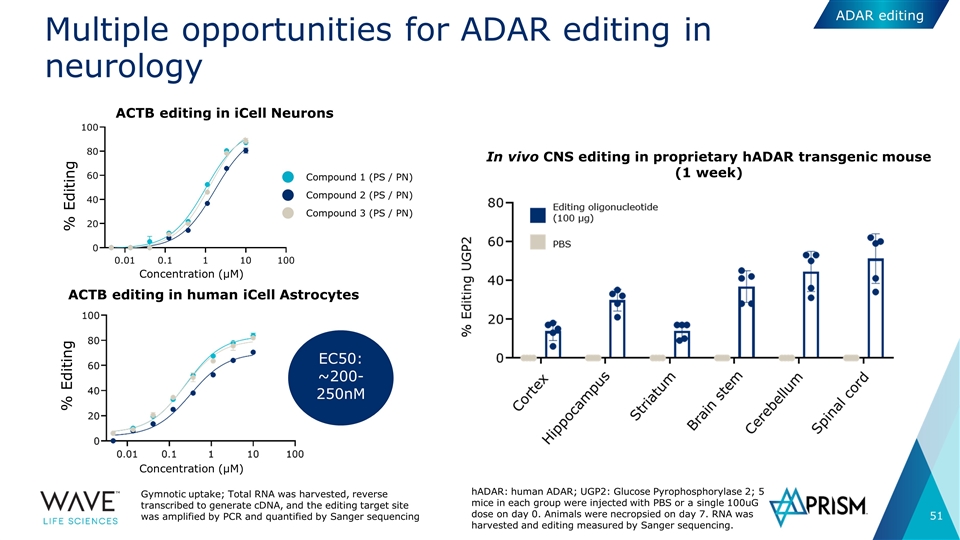
Multiple opportunities for ADAR editing in neurology ACTB editing in iCell Neurons ACTB editing in human iCell Astrocytes Concentration (µM) % Editing % Editing Compound 2 (PS / PN) Compound 1 (PS / PN) Compound 3 (PS / PN) EC50: ~200-250nM Gymnotic uptake; Total RNA was harvested, reverse transcribed to generate cDNA, and the editing target site was amplified by PCR and quantified by Sanger sequencing Concentration (µM) ADAR editing hADAR: human ADAR; UGP2: Glucose Pyrophosphorylase 2; 5 mice in each group were injected with PBS or a single 100uG dose on day 0. Animals were necropsied on day 7. RNA was harvested and editing measured by Sanger sequencing. In vivo CNS editing in proprietary hADAR transgenic mouse (1 week)

Ophthalmology
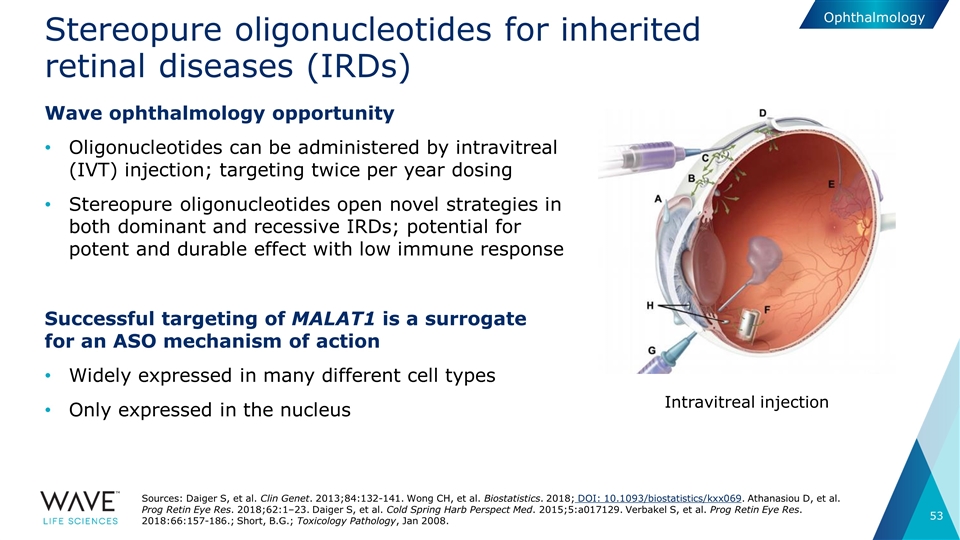
Stereopure oligonucleotides for inherited retinal diseases (IRDs) Wave ophthalmology opportunity Oligonucleotides can be administered by intravitreal (IVT) injection; targeting twice per year dosing Stereopure oligonucleotides open novel strategies in both dominant and recessive IRDs; potential for potent and durable effect with low immune response Successful targeting of MALAT1 is a surrogate for an ASO mechanism of action Widely expressed in many different cell types Only expressed in the nucleus Intravitreal injection Sources: Daiger S, et al. Clin Genet. 2013;84:132-141. Wong CH, et al. Biostatistics. 2018; DOI: 10.1093/biostatistics/kxx069. Athanasiou D, et al. Prog Retin Eye Res. 2018;62:1–23. Daiger S, et al. Cold Spring Harb Perspect Med. 2015;5:a017129. Verbakel S, et al. Prog Retin Eye Res. 2018:66:157-186.; Short, B.G.; Toxicology Pathology, Jan 2008. Ophthalmology
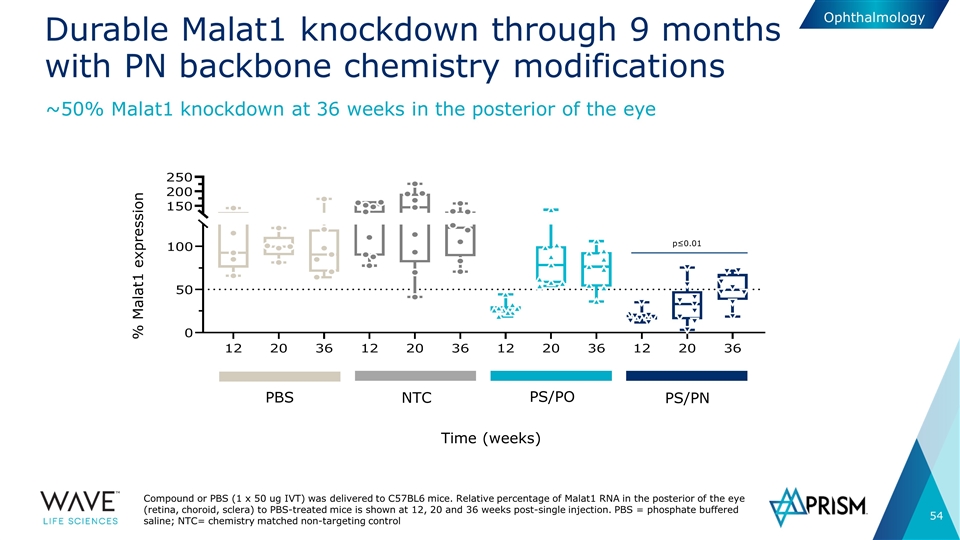
Durable Malat1 knockdown through 9 months with PN backbone chemistry modifications Compound or PBS (1 x 50 ug IVT) was delivered to C57BL6 mice. Relative percentage of Malat1 RNA in the posterior of the eye (retina, choroid, sclera) to PBS-treated mice is shown at 12, 20 and 36 weeks post-single injection. PBS = phosphate buffered saline; NTC= chemistry matched non-targeting control ~50% Malat1 knockdown at 36 weeks in the posterior of the eye PBS NTC PS/PO PS/PN % Malat1 expression Time (weeks) p≤0.01 Ophthalmology
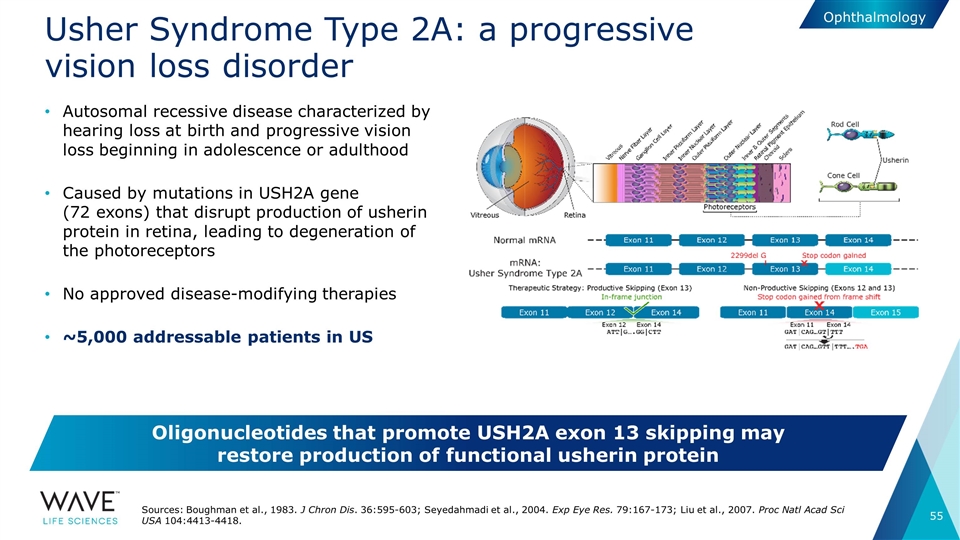
Usher Syndrome Type 2A: a progressive vision loss disorder Autosomal recessive disease characterized by hearing loss at birth and progressive vision loss beginning in adolescence or adulthood Caused by mutations in USH2A gene (72 exons) that disrupt production of usherin protein in retina, leading to degeneration of the photoreceptors No approved disease-modifying therapies ~5,000 addressable patients in US Sources: Boughman et al., 1983. J Chron Dis. 36:595-603; Seyedahmadi et al., 2004. Exp Eye Res. 79:167-173; Liu et al., 2007. Proc Natl Acad Sci USA 104:4413-4418. Oligonucleotides that promote USH2A exon 13 skipping may restore production of functional usherin protein Ophthalmology
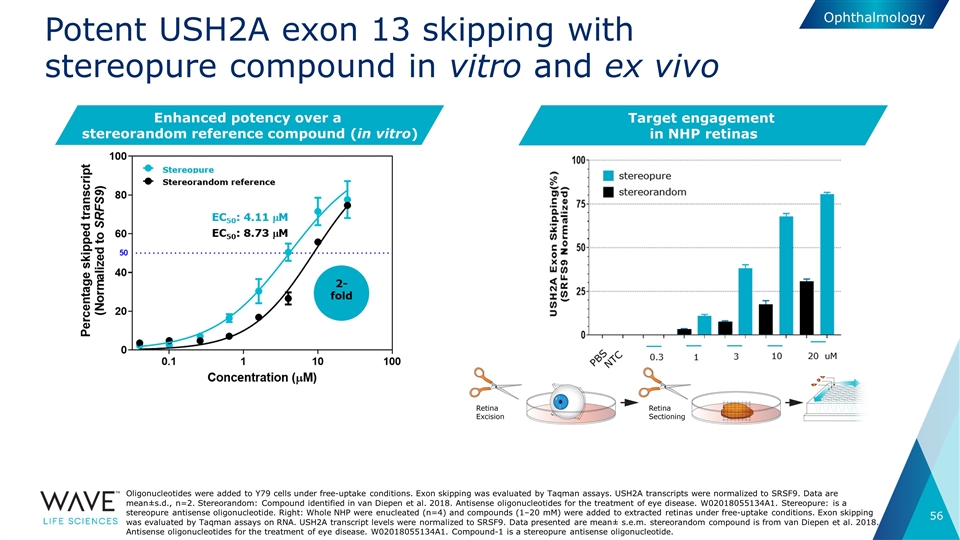
Potent USH2A exon 13 skipping with stereopure compound in vitro and ex vivo Oligonucleotides were added to Y79 cells under free-uptake conditions. Exon skipping was evaluated by Taqman assays. USH2A transcripts were normalized to SRSF9. Data are mean±s.d., n=2. Stereorandom: Compound identified in van Diepen et al. 2018. Antisense oligonucleotides for the treatment of eye disease. W02018055134A1. Stereopure: is a stereopure antisense oligonucleotide. Right: Whole NHP were enucleated (n=4) and compounds (1–20 mM) were added to extracted retinas under free-uptake conditions. Exon skipping was evaluated by Taqman assays on RNA. USH2A transcript levels were normalized to SRSF9. Data presented are mean± s.e.m. stereorandom compound is from van Diepen et al. 2018. Antisense oligonucleotides for the treatment of eye disease. W02018055134A1. Compound-1 is a stereopure antisense oligonucleotide. Enhanced potency over a stereorandom reference compound (in vitro) Ophthalmology Target engagement in NHP retinas
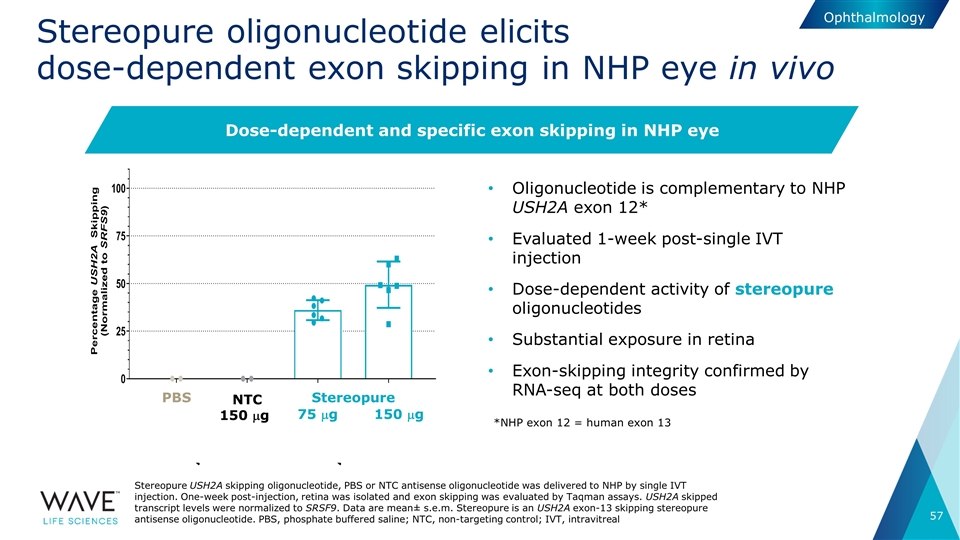
Stereopure oligonucleotide elicits dose-dependent exon skipping in NHP eye in vivo Oligonucleotide is complementary to NHP USH2A exon 12* Evaluated 1-week post-single IVT injection Dose-dependent activity of stereopure oligonucleotides Substantial exposure in retina Exon-skipping integrity confirmed by RNA-seq at both doses NTC 150 mg 75 mg 150 mg PBS Stereopure Stereopure USH2A skipping oligonucleotide, PBS or NTC antisense oligonucleotide was delivered to NHP by single IVT injection. One-week post-injection, retina was isolated and exon skipping was evaluated by Taqman assays. USH2A skipped transcript levels were normalized to SRSF9. Data are mean± s.e.m. Stereopure is an USH2A exon-13 skipping stereopure antisense oligonucleotide. PBS, phosphate buffered saline; NTC, non-targeting control; IVT, intravitreal Dose-dependent and specific exon skipping in NHP eye *NHP exon 12 = human exon 13 Ophthalmology
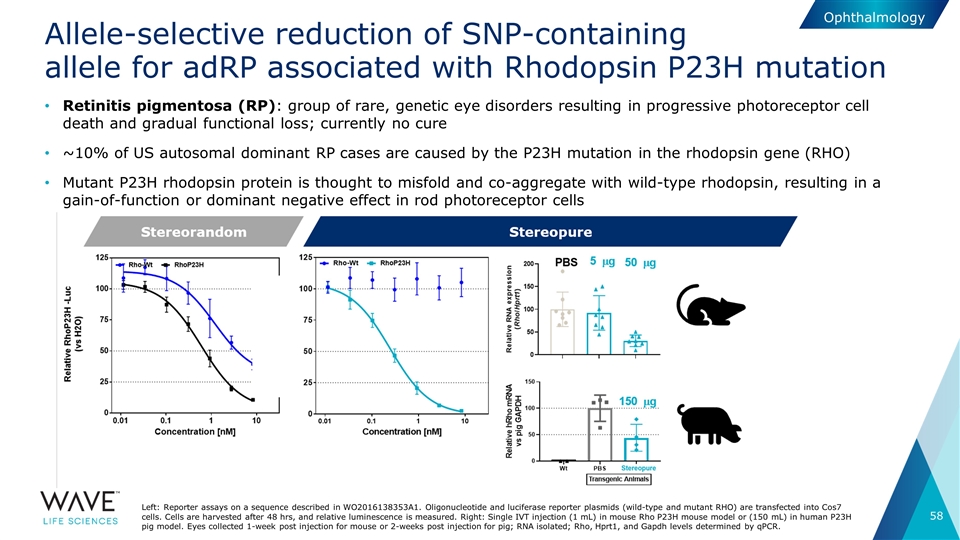
Allele-selective reduction of SNP-containing allele for adRP associated with Rhodopsin P23H mutation Left: Reporter assays on a sequence described in WO2016138353A1. Oligonucleotide and luciferase reporter plasmids (wild-type and mutant RHO) are transfected into Cos7 cells. Cells are harvested after 48 hrs, and relative luminescence is measured. Right: Single IVT injection (1 mL) in mouse Rho P23H mouse model or (150 mL) in human P23H pig model. Eyes collected 1-week post injection for mouse or 2-weeks post injection for pig; RNA isolated; Rho, Hprt1, and Gapdh levels determined by qPCR. Ophthalmology Retinitis pigmentosa (RP): group of rare, genetic eye disorders resulting in progressive photoreceptor cell death and gradual functional loss; currently no cure ~10% of US autosomal dominant RP cases are caused by the P23H mutation in the rhodopsin gene (RHO) Mutant P23H rhodopsin protein is thought to misfold and co-aggregate with wild-type rhodopsin, resulting in a gain-of-function or dominant negative effect in rod photoreceptor cells
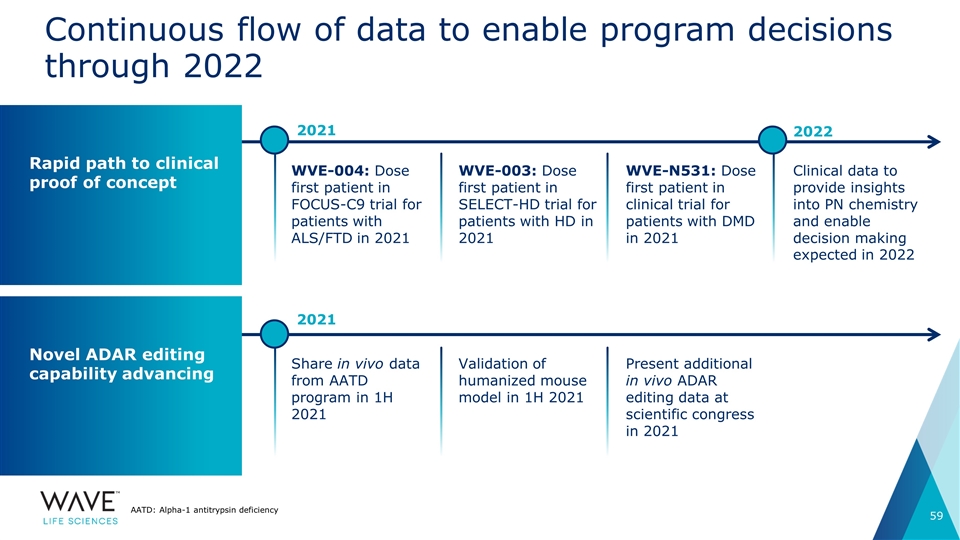
WVE-004: Dose first patient in FOCUS-C9 trial for patients with ALS/FTD in 2021 Continuous flow of data to enable program decisions through 2022 WVE-003: Dose first patient in SELECT-HD trial for patients with HD in 2021 WVE-N531: Dose first patient in clinical trial for patients with DMD in 2021 Clinical data to provide insights into PN chemistry and enable decision making expected in 2022 Rapid path to clinical proof of concept Novel ADAR editing capability advancing Share in vivo data from AATD program in 1H 2021 Validation of humanized mouse model in 1H 2021 Present additional in vivo ADAR editing data at scientific congress in 2021 AATD: Alpha-1 antitrypsin deficiency 2021 2022 2021

Realizing a brighter future for people affected by genetic diseases For more information: Kate Rausch, Investor Relations [email protected] 617.949.4827
Serious News for Serious Traders! Try StreetInsider.com Premium Free!
You May Also Be Interested In
- European Energy A/S: Mitsubishi HC Capital Inc. acquisition of 20% ownership stake in European Energy A/S and election of a new board member
- Alarm.com to Announce 2024 First Quarter Results on May 9, 2024
- Genmab Announces Net Sales of DARZALEX® (daratumumab) for First Quarter of 2024
Create E-mail Alert Related Categories
SEC FilingsSign up for StreetInsider Free!
Receive full access to all new and archived articles, unlimited portfolio tracking, e-mail alerts, custom newswires and RSS feeds - and more!


 Tweet
Tweet Share
Share